Model info
| Transcription factor | CEBPD (GeneCards) | ||||||||
| Model | CEBPD_HUMAN.H11DI.0.C | ||||||||
| Model type | Dinucleotide PWM | ||||||||
| LOGO | 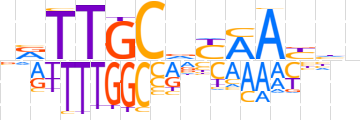 | ||||||||
| LOGO (reverse complement) |  | ||||||||
| Data source | ChIP-Seq | ||||||||
| Model release | HOCOMOCOv11 | ||||||||
| Model length | 13 | ||||||||
| Quality | C | ||||||||
| Motif rank | 0 | ||||||||
| Consensus | nRTTGCnYAAYhn | ||||||||
| Best auROC (human) | 0.809 | ||||||||
| Best auROC (mouse) | |||||||||
| Peak sets in benchmark (human) | 10 | ||||||||
| Peak sets in benchmark (mouse) | |||||||||
| Aligned words | 307 | ||||||||
| TF family | C/EBP-related {1.1.8} | ||||||||
| TF subfamily | C/EBP {1.1.8.1} | ||||||||
| HGNC | HGNC:1835 | ||||||||
| EntrezGene | GeneID:1052 (SSTAR profile) | ||||||||
| UniProt ID | CEBPD_HUMAN | ||||||||
| UniProt AC | P49716 (TFClass) | ||||||||
| Comment | |||||||||
| Downloads | pcm
pwm alignment threshold to P-value grid | ||||||||
| Standard thresholds |
| ||||||||
| GTEx tissue expression atlas | CEBPD expression | ||||||||
| ReMap ChIP-seq dataset list | CEBPD datasets | ||||||||
| Motifs in JASPAR |
PCM
| AA | AC | AG | AT | CA | CC | CG | CT | GA | GC | GG | GT | TA | TC | TG | TT | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 01 | 18.0 | 5.0 | 26.0 | 1.0 | 49.0 | 13.0 | 12.0 | 3.0 | 63.0 | 6.0 | 29.0 | 1.0 | 27.0 | 22.0 | 29.0 | 0.0 |
| 02 | 1.0 | 3.0 | 0.0 | 153.0 | 0.0 | 7.0 | 0.0 | 39.0 | 0.0 | 6.0 | 0.0 | 90.0 | 0.0 | 0.0 | 0.0 | 5.0 |
| 03 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 1.0 | 0.0 | 15.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 5.0 | 281.0 |
| 04 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 1.0 | 4.0 | 3.0 | 1.0 | 267.0 | 26.0 |
| 05 | 0.0 | 3.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 270.0 | 0.0 | 0.0 | 0.0 | 28.0 | 2.0 | 0.0 |
| 06 | 0.0 | 0.0 | 0.0 | 0.0 | 138.0 | 34.0 | 95.0 | 35.0 | 1.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 07 | 15.0 | 94.0 | 4.0 | 26.0 | 0.0 | 27.0 | 0.0 | 8.0 | 4.0 | 53.0 | 3.0 | 35.0 | 0.0 | 25.0 | 0.0 | 10.0 |
| 08 | 5.0 | 14.0 | 0.0 | 0.0 | 175.0 | 22.0 | 2.0 | 0.0 | 2.0 | 4.0 | 0.0 | 1.0 | 32.0 | 47.0 | 0.0 | 0.0 |
| 09 | 210.0 | 3.0 | 1.0 | 0.0 | 80.0 | 3.0 | 0.0 | 4.0 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.0 | 0.0 | 0.0 |
| 10 | 10.0 | 149.0 | 32.0 | 101.0 | 1.0 | 4.0 | 1.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 1.0 | 2.0 | 1.0 | 1.0 |
| 11 | 0.0 | 10.0 | 2.0 | 1.0 | 43.0 | 55.0 | 3.0 | 54.0 | 15.0 | 12.0 | 6.0 | 1.0 | 20.0 | 46.0 | 17.0 | 19.0 |
| 12 | 11.0 | 20.0 | 29.0 | 18.0 | 34.0 | 35.0 | 16.0 | 38.0 | 6.0 | 10.0 | 6.0 | 6.0 | 5.0 | 20.0 | 36.0 | 14.0 |
PWM
| AA | AC | AG | AT | CA | CC | CG | CT | GA | GC | GG | GT | TA | TC | TG | TT | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 01 | -0.053 | -1.285 | 0.309 | -2.658 | 0.936 | -0.371 | -0.449 | -1.752 | 1.186 | -1.113 | 0.416 | -2.658 | 0.346 | 0.144 | 0.416 | -3.992 |
| 02 | -2.658 | -1.752 | -3.992 | 2.07 | -3.992 | -0.967 | -3.992 | 0.71 | -3.992 | -1.113 | -3.992 | 1.541 | -3.992 | -3.992 | -3.992 | -1.285 |
| 03 | -3.992 | -3.992 | -3.992 | -2.658 | -3.992 | -2.658 | -3.992 | -0.231 | -3.992 | -3.992 | -3.992 | -3.992 | -2.658 | -3.992 | -1.285 | 2.677 |
| 04 | -3.992 | -3.992 | -2.658 | -3.992 | -3.992 | -3.992 | -2.658 | -3.992 | -3.992 | -3.992 | -2.658 | -1.491 | -1.752 | -2.658 | 2.626 | 0.309 |
| 05 | -3.992 | -1.752 | -3.992 | -3.992 | -3.992 | -2.658 | -3.992 | -3.992 | -3.992 | 2.637 | -3.992 | -3.992 | -3.992 | 0.382 | -2.106 | -3.992 |
| 06 | -3.992 | -3.992 | -3.992 | -3.992 | 1.967 | 0.574 | 1.595 | 0.602 | -2.658 | -2.658 | -3.992 | -3.992 | -3.992 | -3.992 | -3.992 | -3.992 |
| 07 | -0.231 | 1.584 | -1.491 | 0.309 | -3.992 | 0.346 | -3.992 | -0.84 | -1.491 | 1.014 | -1.752 | 0.602 | -3.992 | 0.27 | -3.992 | -0.625 |
| 08 | -1.285 | -0.299 | -3.992 | -3.992 | 2.204 | 0.144 | -2.106 | -3.992 | -2.106 | -1.491 | -3.992 | -2.658 | 0.514 | 0.895 | -3.992 | -3.992 |
| 09 | 2.386 | -1.752 | -2.658 | -3.992 | 1.423 | -1.752 | -3.992 | -1.491 | -2.658 | -3.992 | -3.992 | -2.658 | -2.658 | -3.992 | -3.992 | -3.992 |
| 10 | -0.625 | 2.043 | 0.514 | 1.656 | -2.658 | -1.491 | -2.658 | -3.992 | -2.658 | -3.992 | -3.992 | -3.992 | -2.658 | -2.106 | -2.658 | -2.658 |
| 11 | -3.992 | -0.625 | -2.106 | -2.658 | 0.806 | 1.051 | -1.752 | 1.033 | -0.231 | -0.449 | -1.113 | -2.658 | 0.05 | 0.873 | -0.109 | 0.0 |
| 12 | -0.533 | 0.05 | 0.416 | -0.053 | 0.574 | 0.602 | -0.168 | 0.684 | -1.113 | -0.625 | -1.113 | -1.113 | -1.285 | 0.05 | 0.63 | -0.299 |