PWMs for MOUSE transcription factors (full)
| Model | LOGO | Transcription factor | Model length | Quality | Model rank | Consensus | Model release | Data source | Best auROC (human) | Best auROC (mouse) | Peak sets in benchmark (human) | Peak sets in benchmark (mouse) | Aligned words | TF family | TF subfamily | MGI | EntrezGene | UniProt ID | UniProt AC |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AHR_MOUSE.H11MO.0.B |  |
MOUSE:Ahr | 9 | B | 0 | dKhGCGTGh | HOCOMOCOv9 | Integrative | 157 | PAS domain factors{1.2.5} | Ahr-like factors{1.2.5.1} | 105043 | 11622 | AHR_MOUSE | P30561 | ||||
| AIRE_MOUSE.H11MO.0.C |  |
MOUSE:Aire | 18 | C | 0 | hnnGGWWnddWWGGdbWh | HOCOMOCOv9 | Integrative | 41 | AIRE{5.3.1} | AIRE{5.3.1.0.1} | 1338803 | 11634 | AIRE_MOUSE | Q9Z0E3 | ||||
| ALX1_MOUSE.H11MO.0.B |  |
MOUSE:Alx1 | 12 | B | 0 | vTRATYGnATTA | HOCOMOCOv9 | Integrative | 33 | Paired-related HD factors{3.1.3} | ALX{3.1.3.1} | 104621 | 216285 | ALX1_MOUSE | Q8C8B0 | ||||
| ANDR_MOUSE.H11MO.0.A |  |
MOUSE:Ar | 16 | A | 0 | RGvACAbTvTGTbCYb | HOCOMOCOv11 | ChIP-Seq | 0.967 | 0.995 | 461 | 57 | 500 | Steroid hormone receptors (NR3){2.1.1} | GR-like receptors (NR3C){2.1.1.1} | 88064 | 11835 | ANDR_MOUSE | P19091 |
| ANDR_MOUSE.H11MO.1.A |  |
MOUSE:Ar | 12 | A | 1 | dbYbTGTbCYbd | HOCOMOCOv11 | ChIP-Seq | 0.897 | 0.929 | 461 | 57 | 500 | Steroid hormone receptors (NR3){2.1.1} | GR-like receptors (NR3C){2.1.1.1} | 88064 | 11835 | ANDR_MOUSE | P19091 |
| AP2A_MOUSE.H11MO.0.A |  |
MOUSE:Tfap2a | 12 | A | 0 | vbdSCCYGRGGC | HOCOMOCOv11 | ChIP-Seq | 0.939 | 0.955 | 13 | 13 | 500 | AP-2{1.3.1} | AP-2alpha{1.3.1.0.1} | 104671 | 21418 | AP2A_MOUSE | P34056 |
| AP2B_MOUSE.H11MO.0.B |  |
MOUSE:Tfap2b | 10 | B | 0 | GSCYGRGGRv | HOCOMOCOv9 | Integrative | 30 | AP-2{1.3.1} | AP-2beta{1.3.1.0.2} | 104672 | 21419 | AP2B_MOUSE | Q61313 | ||||
| AP2C_MOUSE.H11MO.0.A |  |
MOUSE:Tfap2c | 13 | A | 0 | bdSCCYCRGGShn | HOCOMOCOv11 | ChIP-Seq | 0.939 | 0.956 | 18 | 6 | 500 | AP-2{1.3.1} | AP-2gamma{1.3.1.0.3} | 106032 | 21420 | AP2C_MOUSE | Q61312 |
| AP2D_MOUSE.H11MO.0.D |  |
MOUSE:Tfap2d | 14 | D | 0 | nGbCCGRGGCnCGY | HOCOMOCOv9 | Integrative | 5 | AP-2{1.3.1} | AP-2delta{1.3.1.0.4} | 2153466 | 226896 | AP2D_MOUSE | Q91ZK0 | ||||
| ARI3A_MOUSE.H11MO.0.D |  |
MOUSE:Arid3a | 22 | D | 0 | dWTTAAWYvRdWWYAAAWYWWW | HOCOMOCOv9 | Integrative | 44 | ARID-related factors{3.7.1} | ARID3{3.7.1.3} | 1328360 | 13496 | ARI3A_MOUSE | Q62431 | ||||
| ARI5B_MOUSE.H11MO.0.C |  |
MOUSE:Arid5b | 13 | C | 0 | nbYKnRTATTSKd | HOCOMOCOv9 | Integrative | 19 | ARID-related factors{3.7.1} | ARID5{3.7.1.5} | 2175912 | 71371 | ARI5B_MOUSE | Q8BM75 | ||||
| ARNT2_MOUSE.H11MO.0.D |  |
MOUSE:Arnt2 | 12 | D | 0 | RnSGTGGvRSRS | HOCOMOCOv9 | Integrative | 18 | PAS domain factors{1.2.5} | Arnt-like factors{1.2.5.2} | 107188 | 11864 | ARNT2_MOUSE | Q61324 | ||||
| ARNT_MOUSE.H11MO.0.B |  |
MOUSE:Arnt | 8 | B | 0 | bYACGTGh | HOCOMOCOv9 | Integrative | 283 | PAS domain factors{1.2.5} | Arnt-like factors{1.2.5.2} | 88071 | 11863 | ARNT_MOUSE | P53762 | ||||
| ASCL1_MOUSE.H11MO.0.A |  |
MOUSE:Ascl1 | 14 | A | 0 | bhvCASCTGYYbbh | HOCOMOCOv11 | ChIP-Seq | 0.977 | 0.99 | 30 | 25 | 447 | MyoD / ASC-related factors{1.2.2} | Achaete-Scute-like factors{1.2.2.2} | 96919 | 17172 | ASCL1_MOUSE | Q02067 |
| ASCL2_MOUSE.H11MO.0.C |  |
MOUSE:Ascl2 | 9 | C | 0 | vCASCTGCY | HOCOMOCOv11 | ChIP-Seq | 0.594 | 0.92 | 8 | 9 | 505 | MyoD / ASC-related factors{1.2.2} | Achaete-Scute-like factors{1.2.2.2} | 96920 | 17173 | ASCL2_MOUSE | O35885 |
| ATF1_MOUSE.H11MO.0.B |  |
MOUSE:Atf1 | 10 | B | 0 | nTGACGTSMv | HOCOMOCOv9 | Integrative | 43 | CREB-related factors{1.1.7} | CREB-like factors{1.1.7.1} | 1298366 | 11908 | ATF1_MOUSE | P81269 | ||||
| ATF2_MOUSE.H11MO.0.A |  |
MOUSE:Atf2 | 11 | A | 0 | dvTGAbGWMAb | HOCOMOCOv11 | ChIP-Seq | 0.751 | 0.849 | 8 | 4 | 502 | Jun-related factors{1.1.1} | ATF-2-like factors{1.1.1.3} | 109349 | 102641666; 11909 | ATF2_MOUSE | P16951 |
| ATF3_MOUSE.H11MO.0.A |  |
MOUSE:Atf3 | 9 | A | 0 | vTGAGTCAb | HOCOMOCOv11 | ChIP-Seq | 0.905 | 0.887 | 36 | 16 | 500 | Fos-related factors{1.1.2} | ATF-3-like factors{1.1.2.2} | 109384 | 11910 | ATF3_MOUSE | Q60765 |
| ATF4_MOUSE.H11MO.0.A |  |
MOUSE:Atf4 | 12 | A | 0 | dnMTGATGCAAY | HOCOMOCOv11 | ChIP-Seq | 0.998 | 0.955 | 18 | 9 | 500 | ATF-4-related factors{1.1.6} | ATF-4{1.1.6.0.1} | 88096 | 11911 | ATF4_MOUSE | Q06507 |
| ATF7_MOUSE.H11MO.0.D |  |
MOUSE:Atf7 | 11 | D | 0 | vdTGAbGWMAb | HOCOMOCOv11 | ChIP-Seq | 0.596 | 0.773 | 1 | 2 | 500 | Jun-related factors{1.1.1} | ATF-2-like factors{1.1.1.3} | 2443472 | 223922 | ATF7_MOUSE | Q8R0S1 |
| ATOH1_MOUSE.H11MO.0.B |  |
MOUSE:Atoh1 | 9 | B | 0 | RRCAGMTGK | HOCOMOCOv11 | ChIP-Seq | 0.759 | 0.988 | 3 | 4 | 502 | Tal-related factors{1.2.3} | Neurogenin / Atonal-like factors{1.2.3.4} | 104654 | 11921 | ATOH1_MOUSE | P48985 |
| BACH1_MOUSE.H11MO.0.C |  |
MOUSE:Bach1 | 14 | C | 0 | YGCTGAGTCAYGST | HOCOMOCOv9 | Integrative | 8 | Jun-related factors{1.1.1} | NF-E2-like factors{1.1.1.2} | 894680 | 12013 | BACH1_MOUSE | P97302 | ||||
| BACH2_MOUSE.H11MO.0.A |  |
MOUSE:Bach2 | 11 | A | 0 | dSCYSAGTCAY | HOCOMOCOv11 | ChIP-Seq | 0.925 | 0.987 | 10 | 3 | 500 | Jun-related factors{1.1.1} | NF-E2-like factors{1.1.1.2} | 894679 | 12014 | BACH2_MOUSE | P97303 |
| BARX1_MOUSE.H11MO.0.C |  |
MOUSE:Barx1 | 12 | C | 0 | bbWAAdKGYYWh | HOCOMOCOv11 | ChIP-Seq | 0.529 | 0.819 | 2 | 2 | 150 | NK-related factors{3.1.2} | BARX{3.1.2.2} | 103124 | 12022 | BARX1_MOUSE | Q9ER42 |
| BARX2_MOUSE.H11MO.0.D |  |
MOUSE:Barx2 | 10 | D | 0 | hYnWTAATKR | HOCOMOCOv9 | Integrative | 14 | NK-related factors{3.1.2} | BARX{3.1.2.2} | 109617 | 12023 | BARX2_MOUSE | O08686 | ||||
| BATF3_MOUSE.H11MO.0.A |  |
MOUSE:Batf3 | 17 | A | 0 | ndSTTYChnWhTGAbdn | HOCOMOCOv11 | ChIP-Seq | 0.963 | 5 | 500 | B-ATF-related factors{1.1.4} | B-ATF-3{1.1.4.0.3} | 1925491 | 381319 | BATF3_MOUSE | Q9D275 | ||
| BATF_MOUSE.H11MO.0.A |  |
MOUSE:Batf | 18 | A | 0 | dnKhhhvddWTGASTSWb | HOCOMOCOv11 | ChIP-Seq | 0.959 | 0.943 | 10 | 20 | 500 | B-ATF-related factors{1.1.4} | B-ATF{1.1.4.0.1} | 1859147 | 53314 | BATF_MOUSE | O35284 |
| BATF_MOUSE.H11MO.1.A |  |
MOUSE:Batf | 11 | A | 1 | vdWWTGASTMA | HOCOMOCOv11 | ChIP-Seq | 0.914 | 0.93 | 10 | 20 | 500 | B-ATF-related factors{1.1.4} | B-ATF{1.1.4.0.1} | 1859147 | 53314 | BATF_MOUSE | O35284 |
| BCL6_MOUSE.H11MO.0.A |  |
MOUSE:Bcl6 | 13 | A | 0 | hnMWYTCYAGGRW | HOCOMOCOv11 | ChIP-Seq | 0.797 | 0.885 | 27 | 11 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | BCL6 factors{2.3.3.22} | 107187 | 12053 | BCL6_MOUSE | P41183 |
| BHA15_MOUSE.H11MO.0.A |  |
MOUSE:Bhlha15 | 11 | A | 0 | vvRRCAGCTGK | HOCOMOCOv11 | ChIP-Seq | 0.988 | 6 | 500 | Tal-related factors{1.2.3} | Neurogenin / Atonal-like factors{1.2.3.4} | 891976 | 17341 | BHA15_MOUSE | Q9QYC3 | ||
| BHE40_MOUSE.H11MO.0.A |  |
MOUSE:Bhlhe40 | 9 | A | 0 | dCACGTGMv | HOCOMOCOv11 | ChIP-Seq | 0.826 | 0.978 | 10 | 5 | 505 | Hairy-related factors{1.2.4} | Hairy-like factors{1.2.4.1} | 1097714 | 20893 | BHE40_MOUSE | O35185 |
| BHE41_MOUSE.H11MO.0.D |  |
MOUSE:Bhlhe41 | 20 | D | 0 | dCnRnSTCRMGTGMhKRnRn | HOCOMOCOv9 | Integrative | 10 | Hairy-related factors{1.2.4} | Hairy-like factors{1.2.4.1} | 1930704 | 79362 | BHE41_MOUSE | Q99PV5 | ||||
| BMAL1_MOUSE.H11MO.0.A |  |
MOUSE:Arntl | 9 | A | 0 | MCACGTGMb | HOCOMOCOv11 | ChIP-Seq | 0.866 | 0.88 | 18 | 15 | 501 | PAS domain factors{1.2.5} | Arnt-like factors{1.2.5.2} | 1096381 | 11865 | BMAL1_MOUSE | Q9WTL8 |
| BRAC_MOUSE.H11MO.0.B |  |
MOUSE:T | 25 | B | 0 | dKKhvCWbhdddSTGTSWRdKhnKb | HOCOMOCOv11 | ChIP-Seq | 0.704 | 0.814 | 9 | 10 | 435 | Brachyury-related factors{6.5.1} | T (Brachyury){6.5.1.0.1} | 98472 | 20997 | BRAC_MOUSE | P20293 |
| BRAC_MOUSE.H11MO.1.C |  |
MOUSE:T | 13 | C | 1 | dvbSTGTSWRdKb | HOCOMOCOv11 | ChIP-Seq | 0.643 | 0.755 | 9 | 10 | 500 | Brachyury-related factors{6.5.1} | T (Brachyury){6.5.1.0.1} | 98472 | 20997 | BRAC_MOUSE | P20293 |
| BRCA1_MOUSE.H11MO.0.D |  |
MOUSE:Brca1 | 9 | D | 0 | bKKbKGTTG | HOCOMOCOv9 | Integrative | 72 | 104537 | 12189 | BRCA1_MOUSE | P48754 | ||||||
| CDC5L_MOUSE.H11MO.0.D |  |
MOUSE:Cdc5l | 15 | D | 0 | vbGnKdTAAYRWAWb | HOCOMOCOv9 | Integrative | 10 | Myb/SANT domain factors{3.5.1} | Myb-like factors{3.5.1.1} | 1918952 | 71702 | CDC5L_MOUSE | Q6A068 | ||||
| CDX1_MOUSE.H11MO.0.C |  |
MOUSE:Cdx1 | 8 | C | 0 | hTTKAWGd | HOCOMOCOv9 | Integrative | 30 | HOX-related factors{3.1.1} | CDX (Caudal type homeobox){3.1.1.9} | 88360 | 12590 | CDX1_MOUSE | P18111 | ||||
| CDX2_MOUSE.H11MO.0.A |  |
MOUSE:Cdx2 | 12 | A | 0 | bTTTATKRYhbb | HOCOMOCOv11 | ChIP-Seq | 0.916 | 0.99 | 13 | 23 | 500 | HOX-related factors{3.1.1} | CDX (Caudal type homeobox){3.1.1.9} | 88361 | 12591 | CDX2_MOUSE | P43241 |
| CEBPA_MOUSE.H11MO.0.A |  |
MOUSE:Cebpa | 11 | A | 0 | dvTTKhGCAAY | HOCOMOCOv11 | ChIP-Seq | 0.984 | 0.991 | 18 | 153 | 500 | C/EBP-related{1.1.8} | C/EBP{1.1.8.1} | 99480 | 12606 | CEBPA_MOUSE | P53566 |
| CEBPB_MOUSE.H11MO.0.A |  |
MOUSE:Cebpb | 11 | A | 0 | dRTTRhGCAAY | HOCOMOCOv11 | ChIP-Seq | 0.983 | 0.986 | 65 | 158 | 500 | C/EBP-related{1.1.8} | C/EBP{1.1.8.1} | 88373 | 12608 | CEBPB_MOUSE | P28033 |
| CEBPD_MOUSE.H11MO.0.B |  |
MOUSE:Cebpd | 11 | B | 0 | dvTKdbGMAAK | HOCOMOCOv9 | Integrative | 79 | C/EBP-related{1.1.8} | C/EBP{1.1.8.1} | 103573 | 12609 | CEBPD_MOUSE | Q00322 | ||||
| CEBPE_MOUSE.H11MO.0.A |  |
MOUSE:Cebpe | 12 | A | 0 | dTKSGMAAThdb | HOCOMOCOv9 | Integrative | 803 | C/EBP-related{1.1.8} | C/EBP{1.1.8.1} | 103572 | 110794 | CEBPE_MOUSE | Q6PZD9 | ||||
| CEBPG_MOUSE.H11MO.0.B |  |
MOUSE:Cebpg | 12 | B | 0 | ddMTGATGMAAY | HOCOMOCOv11 | ChIP-Seq | 0.724 | 0.97 | 2 | 4 | 493 | C/EBP-related{1.1.8} | C/EBP{1.1.8.1} | 104982 | 12611 | CEBPG_MOUSE | P53568 |
| CEBPZ_MOUSE.H11MO.0.D |  |
MOUSE:Cebpz | 11 | D | 0 | dSTSATTGGCT | HOCOMOCOv9 | Integrative | 28 | 109386 | 12607 | CEBPZ_MOUSE | P53569 | ||||||
| CLOCK_MOUSE.H11MO.0.C |  |
MOUSE:Clock | 14 | C | 0 | SdvnCAMRYGvvvh | HOCOMOCOv11 | ChIP-Seq | 0.658 | 0.735 | 5 | 22 | 253 | PAS domain factors{1.2.5} | Arnt-like factors{1.2.5.2} | 99698 | 12753 | CLOCK_MOUSE | O08785 |
| COE1_MOUSE.H11MO.0.A |  |
MOUSE:Ebf1 | 13 | A | 0 | dbYCCCWRGGGAv | HOCOMOCOv11 | ChIP-Seq | 0.993 | 0.993 | 17 | 55 | 429 | Early B-Cell Factor-related factors{6.1.5} | EBF1 (COE1){6.1.5.0.1} | 95275 | 13591 | COE1_MOUSE | Q07802 |
| COT1_MOUSE.H11MO.0.B |  |
MOUSE:Nr2f1 | 9 | B | 0 | YWRAGGTCA | HOCOMOCOv9 | Integrative | 86 | RXR-related receptors (NR2){2.1.3} | COUP-like receptors (NR2F){2.1.3.5} | 1352451 | 13865 | COT1_MOUSE | Q60632 | ||||
| COT1_MOUSE.H11MO.1.C |  |
MOUSE:Nr2f1 | 12 | C | 1 | SdbYWRAGKTCA | HOCOMOCOv9 | Integrative | 84 | RXR-related receptors (NR2){2.1.3} | COUP-like receptors (NR2F){2.1.3.5} | 1352451 | 13865 | COT1_MOUSE | Q60632 | ||||
| COT2_MOUSE.H11MO.0.A |  |
MOUSE:Nr2f2 | 13 | A | 0 | nhhRAGGTCAvvd | HOCOMOCOv11 | ChIP-Seq | 0.931 | 0.846 | 28 | 3 | 500 | RXR-related receptors (NR2){2.1.3} | COUP-like receptors (NR2F){2.1.3.5} | 1352452 | 11819 | COT2_MOUSE | P43135 |
| COT2_MOUSE.H11MO.1.B |  |
MOUSE:Nr2f2 | 15 | B | 1 | vRGKbCAdRGRKYvv | HOCOMOCOv9 | Integrative | 0.9 | 0.758 | 28 | 3 | 82 | RXR-related receptors (NR2){2.1.3} | COUP-like receptors (NR2F){2.1.3.5} | 1352452 | 11819 | COT2_MOUSE | P43135 |
| COT2_MOUSE.H11MO.2.B |  |
MOUSE:Nr2f2 | 17 | B | 2 | dRdvbhWRAGGKCAvvv | HOCOMOCOv11 | ChIP-Seq | 0.915 | 0.732 | 28 | 3 | 503 | RXR-related receptors (NR2){2.1.3} | COUP-like receptors (NR2F){2.1.3.5} | 1352452 | 11819 | COT2_MOUSE | P43135 |
| CREB1_MOUSE.H11MO.0.A |  |
MOUSE:Creb1 | 11 | A | 0 | ndRTGACGYhW | HOCOMOCOv11 | ChIP-Seq | 0.927 | 0.888 | 63 | 8 | 501 | CREB-related factors{1.1.7} | CREB-like factors{1.1.7.1} | 88494 | 12912 | CREB1_MOUSE | Q01147 |
| CREM_MOUSE.H11MO.0.C |  |
MOUSE:Crem | 11 | C | 0 | SRvTGACGTSA | HOCOMOCOv9 | Integrative | 30 | CREB-related factors{1.1.7} | CREB-like factors{1.1.7.1} | 88495 | 12916 | CREM_MOUSE | P27699 | ||||
| CRX_MOUSE.H11MO.0.A |  |
MOUSE:Crx | 13 | A | 0 | hvRdvGGATTAdv | HOCOMOCOv11 | ChIP-Seq | 0.933 | 8 | 500 | Paired-related HD factors{3.1.3} | OTX{3.1.3.17} | 1194883 | 12951 | CRX_MOUSE | O54751 | ||
| CTCFL_MOUSE.H11MO.0.A |  |
MOUSE:Ctcfl | 18 | A | 0 | vCCdSYAGGKGGCGChvb | HOCOMOCOv11 | ChIP-Seq | 0.97 | 0.996 | 25 | 8 | 495 | More than 3 adjacent zinc finger factors{2.3.3} | CTCF-like factors{2.3.3.50} | 3652571 | 664799 | CTCFL_MOUSE | A2APF3 |
| CTCF_MOUSE.H11MO.0.A |  |
MOUSE:Ctcf | 20 | A | 0 | bbdCCRSYAGGKGGCGSbvb | HOCOMOCOv11 | ChIP-Seq | 0.998 | 0.999 | 502 | 263 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | CTCF-like factors{2.3.3.50} | 109447 | 13018 | CTCF_MOUSE | Q61164 |
| CUX1_MOUSE.H11MO.0.C |  |
MOUSE:Cux1 | 14 | C | 0 | vbRvndATYRRTbb | HOCOMOCOv9 | Integrative | 97 | HD-CUT factors{3.1.9} | CUX{3.1.9.2} | 88568 | 13047 | CUX1_MOUSE | P53564 | ||||
| CUX2_MOUSE.H11MO.0.C |  |
MOUSE:Cux2 | 11 | C | 0 | hRRTCAATvvv | HOCOMOCOv11 | ChIP-Seq | 0.867 | 5 | 496 | HD-CUT factors{3.1.9} | CUX{3.1.9.2} | 107321 | 13048 | CUX2_MOUSE | P70298 | ||
| CXXC1_MOUSE.H11MO.0.D |  |
MOUSE:Cxxc1 | 7 | D | 0 | YGKTKKY | HOCOMOCOv9 | Integrative | 63 | 1921572 | 74322 | CXXC1_MOUSE | Q9CWW7 | ||||||
| DBP_MOUSE.H11MO.0.B |  |
MOUSE:Dbp | 11 | B | 0 | vTKRYGTAAbv | HOCOMOCOv9 | Integrative | 26 | C/EBP-related{1.1.8} | PAR factors{1.1.8.2} | 94866 | 13170 | DBP_MOUSE | Q60925 | ||||
| DDIT3_MOUSE.H11MO.0.C |  |
MOUSE:Ddit3 | 10 | C | 0 | MTGATGMAAb | HOCOMOCOv11 | ChIP-Seq | 0.628 | 0.91 | 3 | 6 | 500 | C/EBP-related{1.1.8} | C/EBP{1.1.8.1} | 109247 | 13198 | DDIT3_MOUSE | P35639 |
| DLX2_MOUSE.H11MO.0.D |  |
MOUSE:Dlx2 | 8 | D | 0 | ATAATTAn | HOCOMOCOv9 | Integrative | 6 | NK-related factors{3.1.2} | DLX{3.1.2.5} | 94902 | 13392 | DLX2_MOUSE | P40764 | ||||
| DLX3_MOUSE.H11MO.0.C |  |
MOUSE:Dlx3 | 10 | C | 0 | RMTAATTRvh | HOCOMOCOv9 | Integrative | 35 | NK-related factors{3.1.2} | DLX{3.1.2.5} | 94903 | 13393 | DLX3_MOUSE | Q64205 | ||||
| DLX5_MOUSE.H11MO.0.C |  |
MOUSE:Dlx5 | 11 | C | 0 | WRATTRYWKbb | HOCOMOCOv11 | ChIP-Seq | 0.909 | 2 | 465 | NK-related factors{3.1.2} | DLX{3.1.2.5} | 101926 | 13395 | DLX5_MOUSE | P70396 | ||
| DMRT1_MOUSE.H11MO.0.C |  |
MOUSE:Dmrt1 | 17 | C | 0 | hdGhhACAAWGTWdChv | HOCOMOCOv11 | ChIP-Seq | 0.552 | 0.987 | 2 | 4 | 500 | DMRT{2.5.1} | DMRT1{2.5.1.0.1} | 1354733 | 50796 | DMRT1_MOUSE | Q9QZ59 |
| DMRTB_MOUSE.H11MO.0.C |  |
MOUSE:Dmrtb1 | 16 | C | 0 | bdGhWACWTTGTddCh | HOCOMOCOv11 | ChIP-Seq | 0.899 | 3 | 500 | DMRT{2.5.1} | DMRTB1{2.5.1.0.6} | 1927125 | 56296 | DMRTB_MOUSE | A2A9I7 | ||
| E2F1_MOUSE.H11MO.0.A |  |
MOUSE:E2f1 | 12 | A | 0 | vdRGGCGGGRvv | HOCOMOCOv11 | ChIP-Seq | 0.892 | 0.974 | 80 | 21 | 500 | E2F-related factors{3.3.2} | E2F{3.3.2.1} | 101941 | 13555 | E2F1_MOUSE | Q61501 |
| E2F2_MOUSE.H11MO.0.B |  |
MOUSE:E2f2 | 10 | B | 0 | RGCGCGMAWC | HOCOMOCOv9 | Integrative | 8 | E2F-related factors{3.3.2} | E2F{3.3.2.1} | 1096341 | 242705 | E2F2_MOUSE | P56931 | ||||
| E2F3_MOUSE.H11MO.0.A |  |
MOUSE:E2f3 | 10 | A | 0 | dSGCGGGARv | HOCOMOCOv11 | ChIP-Seq | 0.813 | 0.971 | 3 | 22 | 498 | E2F-related factors{3.3.2} | E2F{3.3.2.1} | 1096340 | 13557 | E2F3_MOUSE | O35261 |
| E2F4_MOUSE.H11MO.0.A |  |
MOUSE:E2f4 | 14 | A | 0 | vRddGGCGGGRvnn | HOCOMOCOv11 | ChIP-Seq | 0.949 | 0.977 | 15 | 37 | 500 | E2F-related factors{3.3.2} | E2F{3.3.2.1} | 103012 | 104394 | E2F4_MOUSE | Q8R0K9 |
| E2F4_MOUSE.H11MO.1.A |  |
MOUSE:E2f4 | 14 | A | 1 | SGCGGGRRddbbnd | HOCOMOCOv11 | ChIP-Seq | 0.924 | 0.97 | 15 | 37 | 500 | E2F-related factors{3.3.2} | E2F{3.3.2.1} | 103012 | 104394 | E2F4_MOUSE | Q8R0K9 |
| E2F5_MOUSE.H11MO.0.B |  |
MOUSE:E2f5 | 10 | B | 0 | nGCGCCAAAh | HOCOMOCOv9 | Integrative | 12 | E2F-related factors{3.3.2} | E2F{3.3.2.1} | 105091 | 13559 | E2F5_MOUSE | Q61502 | ||||
| E2F6_MOUSE.H11MO.0.A |  |
MOUSE:E2f6 | 12 | A | 0 | ndRGCGGGARvv | HOCOMOCOv9 | Integrative | 2029 | E2F-related factors{3.3.2} | E2F{3.3.2.1} | 1354159 | 50496 | E2F6_MOUSE | O54917 | ||||
| E2F7_MOUSE.H11MO.0.C |  |
MOUSE:E2f7 | 13 | C | 0 | vdRGCGGGARvdv | HOCOMOCOv11 | ChIP-Seq | 0.846 | 16 | 500 | E2F-related factors{3.3.2} | E2F{3.3.2.1} | 1289147 | 52679 | E2F7_MOUSE | Q6S7F2 | ||
| E4F1_MOUSE.H11MO.0.D |  |
MOUSE:E4f1 | 9 | D | 0 | nGTGACGTM | HOCOMOCOv9 | Integrative | 9 | Factors with multiple dispersed zinc fingers{2.3.4} | unclassified{2.3.4.0} | 109530 | 13560 | E4F1_MOUSE | Q8CCE9 | ||||
| EGR1_MOUSE.H11MO.0.A |  |
MOUSE:Egr1 | 14 | A | 0 | vbGYGKGGGYGGdd | HOCOMOCOv11 | ChIP-Seq | 0.988 | 0.985 | 36 | 12 | 492 | Three-zinc finger Krüppel-related factors{2.3.1} | EGR factors{2.3.1.3} | 95295 | 13653 | EGR1_MOUSE | P08046 |
| EGR2_MOUSE.H11MO.0.A |  |
MOUSE:Egr2 | 14 | A | 0 | bShGKGGGhGKddv | HOCOMOCOv11 | ChIP-Seq | 0.948 | 0.995 | 4 | 25 | 506 | Three-zinc finger Krüppel-related factors{2.3.1} | EGR factors{2.3.1.3} | 95296 | 13654 | EGR2_MOUSE | P08152 |
| EGR2_MOUSE.H11MO.1.A |  |
MOUSE:Egr2 | 18 | A | 1 | vdGhGKGGGhGKvdvddd | HOCOMOCOv11 | ChIP-Seq | 0.942 | 0.993 | 4 | 25 | 508 | Three-zinc finger Krüppel-related factors{2.3.1} | EGR factors{2.3.1.3} | 95296 | 13654 | EGR2_MOUSE | P08152 |
| EGR3_MOUSE.H11MO.0.D |  |
MOUSE:Egr3 | 11 | D | 0 | WRMGTGGGTGb | HOCOMOCOv9 | Integrative | 8 | Three-zinc finger Krüppel-related factors{2.3.1} | EGR factors{2.3.1.3} | 1306780 | 13655 | EGR3_MOUSE | P43300 | ||||
| EGR4_MOUSE.H11MO.0.D |  |
MOUSE:Egr4 | 11 | D | 0 | GGSGGYAGGGM | HOCOMOCOv9 | Integrative | 6 | Three-zinc finger Krüppel-related factors{2.3.1} | EGR factors{2.3.1.3} | 99252 | 13656 | EGR4_MOUSE | Q9WUF2 | ||||
| EHF_MOUSE.H11MO.0.B |  |
MOUSE:Ehf | 15 | B | 0 | ndAbSAGGAAGYdvv | HOCOMOCOv11 | ChIP-Seq | 0.966 | 0.654 | 5 | 3 | 500 | Ets-related factors{3.5.2} | EHF-like factors{3.5.2.4} | 1270840 | 13661 | EHF_MOUSE | O70273 |
| ELF1_MOUSE.H11MO.0.A |  |
MOUSE:Elf1 | 14 | A | 0 | vvRvSCGGAAGYSv | HOCOMOCOv11 | ChIP-Seq | 0.992 | 0.975 | 43 | 5 | 500 | Ets-related factors{3.5.2} | Elf-1-like factors{3.5.2.3} | 107180 | 13709 | ELF1_MOUSE | Q60775 |
| ELF2_MOUSE.H11MO.0.C |  |
MOUSE:Elf2 | 15 | C | 0 | YRnCAGGAAGbGnnK | HOCOMOCOv9 | Integrative | 9 | Ets-related factors{3.5.2} | Elf-1-like factors{3.5.2.3} | 1916507 | 69257 | ELF2_MOUSE | Q9JHC9 | ||||
| ELF3_MOUSE.H11MO.0.B |  |
MOUSE:Elf3 | 14 | B | 0 | vdAbSAGGAARYdv | HOCOMOCOv11 | ChIP-Seq | 0.979 | 11 | 500 | Ets-related factors{3.5.2} | EHF-like factors{3.5.2.4} | 1101781 | 13710 | ELF3_MOUSE | Q3UPW2 | ||
| ELF5_MOUSE.H11MO.0.A |  |
MOUSE:Elf5 | 17 | A | 0 | RRRARRAGGAARddvnd | HOCOMOCOv11 | ChIP-Seq | 0.845 | 0.991 | 3 | 12 | 501 | Ets-related factors{3.5.2} | EHF-like factors{3.5.2.4} | 1335079 | 13711 | ELF5_MOUSE | Q8VDK3 |
| ELK1_MOUSE.H11MO.0.B |  |
MOUSE:Elk1 | 11 | B | 0 | vSCGGAAGYdv | HOCOMOCOv11 | ChIP-Seq | 0.975 | 0.715 | 4 | 7 | 501 | Ets-related factors{3.5.2} | Elk-like factors{3.5.2.2} | 101833 | 13712 | ELK1_MOUSE | P41969 |
| ELK3_MOUSE.H11MO.0.D |  |
MOUSE:Elk3 | 12 | D | 0 | vnCMGGAWGTGC | HOCOMOCOv9 | Integrative | 7 | Ets-related factors{3.5.2} | Elk-like factors{3.5.2.2} | 101762 | 13713 | ELK3_MOUSE | P41971 | ||||
| ELK4_MOUSE.H11MO.0.B |  |
MOUSE:Elk4 | 13 | B | 0 | vvvSMGGAARbvv | HOCOMOCOv11 | ChIP-Seq | 0.955 | 0.647 | 13 | 4 | 175 | Ets-related factors{3.5.2} | Elk-like factors{3.5.2.2} | 102853 | 13714 | ELK4_MOUSE | P41158 |
| EPAS1_MOUSE.H11MO.0.C |  |
MOUSE:Epas1 | 9 | C | 0 | vdACGTGhh | HOCOMOCOv11 | ChIP-Seq | 0.838 | 24 | 500 | PAS domain factors{1.2.5} | Ahr-like factors{1.2.5.1} | 109169 | 13819 | EPAS1_MOUSE | P97481 | ||
| ERG_MOUSE.H11MO.0.A |  |
MOUSE:Erg | 14 | A | 0 | vvvSMGGAARhdvv | HOCOMOCOv11 | ChIP-Seq | 0.963 | 0.976 | 79 | 16 | 500 | Ets-related factors{3.5.2} | Ets-like factors{3.5.2.1} | 95415 | 13876 | ERG_MOUSE | P81270 |
| ERR1_MOUSE.H11MO.0.A |  |
MOUSE:Esrra | 12 | A | 0 | ddnYCAAGGTCA | HOCOMOCOv11 | ChIP-Seq | 0.95 | 0.982 | 31 | 3 | 500 | Steroid hormone receptors (NR3){2.1.1} | ER-like receptors (NR3A&B){2.1.1.2} | 1346831 | 26379 | ERR1_MOUSE | O08580 |
| ERR2_MOUSE.H11MO.0.A |  |
MOUSE:Esrrb | 17 | A | 0 | dhdbYCAAGGTCAhhbh | HOCOMOCOv11 | ChIP-Seq | 0.966 | 14 | 502 | Steroid hormone receptors (NR3){2.1.1} | ER-like receptors (NR3A&B){2.1.1.2} | 1346832 | 26380 | ERR2_MOUSE | Q61539 | ||
| ERR3_MOUSE.H11MO.0.C |  |
MOUSE:Esrrg | 10 | C | 0 | bSAAGGWSAb | HOCOMOCOv11 | ChIP-Seq | 0.849 | 2 | 242 | Steroid hormone receptors (NR3){2.1.1} | ER-like receptors (NR3A&B){2.1.1.2} | 1347056 | 26381 | ERR3_MOUSE | P62509 | ||
| ESR1_MOUSE.H11MO.0.A |  |
MOUSE:Esr1 | 15 | A | 0 | RKGWCASCvTGhYCY | HOCOMOCOv11 | ChIP-Seq | 0.966 | 0.993 | 432 | 58 | 500 | Steroid hormone receptors (NR3){2.1.1} | ER-like receptors (NR3A&B){2.1.1.2} | 1352467 | 13982 | ESR1_MOUSE | P19785 |
| ESR1_MOUSE.H11MO.1.A |  |
MOUSE:Esr1 | 10 | A | 1 | hhRGGTCAbv | HOCOMOCOv11 | ChIP-Seq | 0.935 | 0.957 | 432 | 58 | 500 | Steroid hormone receptors (NR3){2.1.1} | ER-like receptors (NR3A&B){2.1.1.2} | 1352467 | 13982 | ESR1_MOUSE | P19785 |
| ESR2_MOUSE.H11MO.0.A |  |
MOUSE:Esr2 | 15 | A | 0 | RGGbCRbCbYGhCCY | HOCOMOCOv11 | ChIP-Seq | 0.944 | 0.941 | 11 | 7 | 497 | Steroid hormone receptors (NR3){2.1.1} | ER-like receptors (NR3A&B){2.1.1.2} | 109392 | 13983 | ESR2_MOUSE | O08537 |
| ESR2_MOUSE.H11MO.1.A |  |
MOUSE:Esr2 | 8 | A | 1 | RGGKCASn | HOCOMOCOv11 | ChIP-Seq | 0.859 | 0.85 | 11 | 7 | 501 | Steroid hormone receptors (NR3){2.1.1} | ER-like receptors (NR3A&B){2.1.1.2} | 109392 | 13983 | ESR2_MOUSE | O08537 |
| ETS1_MOUSE.H11MO.0.A |  |
MOUSE:Ets1 | 14 | A | 0 | vvvRSMGGAAGYSS | HOCOMOCOv11 | ChIP-Seq | 0.953 | 0.964 | 45 | 33 | 500 | Ets-related factors{3.5.2} | Ets-like factors{3.5.2.1} | 95455 | 23871 | ETS1_MOUSE | P27577 |
| ETS2_MOUSE.H11MO.0.A |  |
MOUSE:Ets2 | 13 | A | 0 | ddvdRGAARvRdv | HOCOMOCOv11 | ChIP-Seq | 0.836 | 12 | 500 | 95456 | 23872 | ETS2_MOUSE | P15037 | ||||
| ETV2_MOUSE.H11MO.0.A |  |
MOUSE:Etv2 | 16 | A | 0 | vvvRRCAGGAARYvSv | HOCOMOCOv11 | ChIP-Seq | 0.981 | 6 | 230 | Ets-related factors{3.5.2} | Ets-like factors{3.5.2.1} | 99253 | 14008 | ETV2_MOUSE | P41163 | ||
| ETV4_MOUSE.H11MO.0.B |  |
MOUSE:Etv4 | 8 | B | 0 | vMGGAAGY | HOCOMOCOv9 | Integrative | 47 | Ets-related factors{3.5.2} | Elk-like factors{3.5.2.2} | 99423 | 18612 | ETV4_MOUSE | P28322 | ||||
| ETV5_MOUSE.H11MO.0.D |  |
MOUSE:Etv5 | 14 | D | 0 | vddvAGGRARddvv | HOCOMOCOv11 | ChIP-Seq | 0.867 | 3 | 500 | Ets-related factors{3.5.2} | Elk-like factors{3.5.2.2} | 1096867 | 104156 | ETV5_MOUSE | Q9CXC9 | ||
| ETV6_MOUSE.H11MO.0.C |  |
MOUSE:Etv6 | 9 | C | 0 | dSAGGAARb | HOCOMOCOv11 | ChIP-Seq | 0.853 | 3 | 500 | Ets-related factors{3.5.2} | ETV6-like factors{3.5.2.6} | 109336 | 14011 | ETV6_MOUSE | P97360 | ||
| EVI1_MOUSE.H11MO.0.B |  |
MOUSE:Mecom | 16 | B | 0 | dWGAYAAGATAAnMbd | HOCOMOCOv9 | Integrative | 141 | Factors with multiple dispersed zinc fingers{2.3.4} | Evi-1-like factors{2.3.4.14} | 95457 | EVI1_MOUSE | P14404 | |||||
| FEV_MOUSE.H11MO.0.B |  |
MOUSE:Fev | 10 | B | 0 | vvvRGARRhS | HOCOMOCOv11 | ChIP-Seq | 0.753 | 0.887 | 2 | 4 | 498 | Ets-related factors{3.5.2} | Ets-like factors{3.5.2.1} | 2449712 | 260298 | FEV_MOUSE | Q8QZW2 |
| FLI1_MOUSE.H11MO.0.A |  |
MOUSE:Fli1 | 14 | A | 0 | vvvdSMGGAAGbvv | HOCOMOCOv11 | ChIP-Seq | 0.953 | 0.958 | 50 | 32 | 500 | Ets-related factors{3.5.2} | Ets-like factors{3.5.2.1} | 95554 | 14247 | FLI1_MOUSE | P26323 |
| FLI1_MOUSE.H11MO.1.A |  |
MOUSE:Fli1 | 19 | A | 1 | vvvvvRSAGGAAGbvvvvv | HOCOMOCOv11 | ChIP-Seq | 0.983 | 0.939 | 50 | 32 | 515 | Ets-related factors{3.5.2} | Ets-like factors{3.5.2.1} | 95554 | 14247 | FLI1_MOUSE | P26323 |
| FOSB_MOUSE.H11MO.0.A |  |
MOUSE:Fosb | 9 | A | 0 | vTGAGTCAb | HOCOMOCOv11 | ChIP-Seq | 0.975 | 0.925 | 2 | 27 | 500 | Fos-related factors{1.1.2} | Fos factors{1.1.2.1} | 95575 | 14282 | FOSB_MOUSE | P13346 |
| FOSL1_MOUSE.H11MO.0.A |  |
MOUSE:Fosl1 | 12 | A | 0 | ddvTGAGTCAbh | HOCOMOCOv11 | ChIP-Seq | 0.988 | 0.952 | 21 | 2 | 500 | Fos-related factors{1.1.2} | Fos factors{1.1.2.1} | 107179 | 14283 | FOSL1_MOUSE | P48755 |
| FOSL2_MOUSE.H11MO.0.A |  |
MOUSE:Fosl2 | 11 | A | 0 | dvTGAGTCAbh | HOCOMOCOv11 | ChIP-Seq | 0.996 | 0.977 | 27 | 5 | 500 | Fos-related factors{1.1.2} | Fos factors{1.1.2.1} | 102858 | 14284 | FOSL2_MOUSE | P47930 |
| FOS_MOUSE.H11MO.0.A |  |
MOUSE:Fos | 12 | A | 0 | vdRTGAGTCAYh | HOCOMOCOv11 | ChIP-Seq | 0.993 | 0.927 | 38 | 50 | 500 | Fos-related factors{1.1.2} | Fos factors{1.1.2.1} | 95574 | 14281 | FOS_MOUSE | P01101 |
| FOXA1_MOUSE.H11MO.0.A | 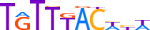 |
MOUSE:Foxa1 | 10 | A | 0 | TGTTTACWYW | HOCOMOCOv11 | ChIP-Seq | 0.98 | 0.978 | 199 | 28 | 501 | Forkhead box (FOX) factors{3.3.1} | FOXA{3.3.1.1} | 1347472 | 15375 | FOXA1_MOUSE | P35582 |
| FOXA2_MOUSE.H11MO.0.A |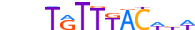  |
MOUSE:Foxa2 | 13 | A | 0 | bnhTGTTTACWbW | HOCOMOCOv11 | ChIP-Seq | 0.977 | 0.992 | 22 | 34 | 500 | Forkhead box (FOX) factors{3.3.1} | FOXA{3.3.1.1} | 1347476 | 15376 | FOXA2_MOUSE | P35583 |
| FOXA3_MOUSE.H11MO.0.A |  |
MOUSE:Foxa3 | 13 | A | 0 | bhTGTTTACWbWd | HOCOMOCOv11 | ChIP-Seq | 0.988 | 6 | 500 | Forkhead box (FOX) factors{3.3.1} | FOXA{3.3.1.1} | 1347477 | 15377 | FOXA3_MOUSE | P35584 | ||
| FOXC1_MOUSE.H11MO.0.C |  |
MOUSE:Foxc1 | 15 | C | 0 | bnhTGTTTACWTAvS | HOCOMOCOv9 | Integrative | 26 | Forkhead box (FOX) factors{3.3.1} | FOXC{3.3.1.3} | 1347466 | 17300 | FOXC1_MOUSE | Q61572 | ||||
| FOXC2_MOUSE.H11MO.0.D |  |
MOUSE:Foxc2 | 15 | D | 0 | KhTTGTTdTGYCAGd | HOCOMOCOv9 | Integrative | 5 | Forkhead box (FOX) factors{3.3.1} | FOXC{3.3.1.3} | 1347481 | 14234 | FOXC2_MOUSE | Q61850 | ||||
| FOXD1_MOUSE.H11MO.0.D |  |
MOUSE:Foxd1 | 15 | D | 0 | nhhTSTTTAChWhvR | HOCOMOCOv9 | Integrative | 22 | 1347463 | 15229 | FOXD1_MOUSE | Q61345 | ||||||
| FOXD3_MOUSE.H11MO.0.C | 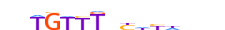 |
MOUSE:Foxd3 | 15 | C | 0 | bbTGTTTnYhYddbh | HOCOMOCOv11 | ChIP-Seq | 0.786 | 11 | 465 | Forkhead box (FOX) factors{3.3.1} | FOXD{3.3.1.4} | 1347473 | 15221 | FOXD3_MOUSE | Q61060 | ||
| FOXF1_MOUSE.H11MO.0.D |  |
MOUSE:Foxf1 | 11 | D | 0 | nTGTTTATbWW | HOCOMOCOv9 | Integrative | 8 | Forkhead box (FOX) factors{3.3.1} | FOXF{3.3.1.6} | 1347470 | 15227 | FOXF1_MOUSE | Q61080 | ||||
| FOXF2_MOUSE.H11MO.0.D |  |
MOUSE:Foxf2 | 11 | D | 0 | TGTTTAChTWd | HOCOMOCOv9 | Integrative | 26 | Forkhead box (FOX) factors{3.3.1} | FOXF{3.3.1.6} | 1347479 | 14238 | FOXF2_MOUSE | O54743 | ||||
| FOXI1_MOUSE.H11MO.0.B |  |
MOUSE:Foxi1 | 12 | B | 0 | hWbTGATTGGYb | HOCOMOCOv9 | Integrative | 210 | Forkhead box (FOX) factors{3.3.1} | FOXI{3.3.1.9} | 1096329 | 14233 | FOXI1_MOUSE | Q922I5 | ||||
| FOXJ2_MOUSE.H11MO.0.C |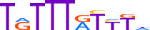  |
MOUSE:Foxj2 | 10 | C | 0 | TRTTTATYTd | HOCOMOCOv9 | Integrative | 41 | Forkhead box (FOX) factors{3.3.1} | FOXJ{3.3.1.10} | 1926805 | 60611 | FOXJ2_MOUSE | Q9ES18 | ||||
| FOXJ3_MOUSE.H11MO.0.A |  |
MOUSE:Foxj3 | 13 | A | 0 | dTGTTTATKKTTd | HOCOMOCOv9 | Integrative | 550 | Forkhead box (FOX) factors{3.3.1} | FOXJ{3.3.1.10} | 2443432 | 230700 | FOXJ3_MOUSE | Q8BUR3 | ||||
| FOXJ3_MOUSE.H11MO.1.B |  |
MOUSE:Foxj3 | 13 | B | 1 | WKKTTWWTGTTTW | HOCOMOCOv9 | Integrative | 499 | Forkhead box (FOX) factors{3.3.1} | FOXJ{3.3.1.10} | 2443432 | 230700 | FOXJ3_MOUSE | Q8BUR3 | ||||
| FOXK1_MOUSE.H11MO.0.A |  |
MOUSE:Foxk1 | 11 | A | 0 | bbTGTTKMYnb | HOCOMOCOv11 | ChIP-Seq | 0.846 | 0.891 | 11 | 4 | 500 | Forkhead box (FOX) factors{3.3.1} | FOXK{3.3.1.11} | 1347488 | 17425 | FOXK1_MOUSE | P42128 |
| FOXL2_MOUSE.H11MO.0.C |  |
MOUSE:Foxl2 | 11 | C | 0 | TGTTTWYhWWh | HOCOMOCOv11 | ChIP-Seq | 0.926 | 2 | 501 | Forkhead box (FOX) factors{3.3.1} | FOXL{3.3.1.12} | 1349428 | 26927 | FOXL2_MOUSE | O88470 | ||
| FOXM1_MOUSE.H11MO.0.B |  |
MOUSE:Foxm1 | 12 | B | 0 | TRTTTRYWbWbn | HOCOMOCOv11 | ChIP-Seq | 0.882 | 66 | 500 | Forkhead box (FOX) factors{3.3.1} | FOXM{3.3.1.13} | 1347487 | FOXM1_MOUSE | O08696 | |||
| FOXO1_MOUSE.H11MO.0.A |  |
MOUSE:Foxo1 | 10 | A | 0 | bbnTGTTTWC | HOCOMOCOv11 | ChIP-Seq | 0.881 | 0.934 | 9 | 21 | 500 | Forkhead box (FOX) factors{3.3.1} | FOXO{3.3.1.15} | 1890077 | 56458 | FOXO1_MOUSE | Q9R1E0 |
| FOXO3_MOUSE.H11MO.0.B |  |
MOUSE:Foxo3 | 10 | B | 0 | bbTGTTTACh | HOCOMOCOv11 | ChIP-Seq | 0.668 | 0.948 | 2 | 7 | 500 | Forkhead box (FOX) factors{3.3.1} | FOXO{3.3.1.15} | 1890081 | 56484 | FOXO3_MOUSE | Q9WVH4 |
| FOXO4_MOUSE.H11MO.0.C |  |
MOUSE:Foxo4 | 9 | C | 0 | bTGTTTAbK | HOCOMOCOv9 | Integrative | 28 | Forkhead box (FOX) factors{3.3.1} | FOXO{3.3.1.15} | 1891915 | 54601 | FOXO4_MOUSE | Q9WVH3 | ||||
| FOXP2_MOUSE.H11MO.0.A |  |
MOUSE:Foxp2 | 9 | A | 0 | nTGTTTMYn | HOCOMOCOv9 | Integrative | 2075 | Forkhead box (FOX) factors{3.3.1} | FOXP{3.3.1.16} | 2148705 | 114142 | FOXP2_MOUSE | P58463 | ||||
| FOXP3_MOUSE.H11MO.0.D |  |
MOUSE:Foxp3 | 9 | D | 0 | WATKTGWTT | HOCOMOCOv9 | Integrative | 9 | Forkhead box (FOX) factors{3.3.1} | FOXP{3.3.1.16} | 1891436 | 20371 | FOXP3_MOUSE | Q99JB6 | ||||
| FOXQ1_MOUSE.H11MO.0.C |  |
MOUSE:Foxq1 | 12 | C | 0 | nRTTGTTTATKT | HOCOMOCOv9 | Integrative | 13 | Forkhead box (FOX) factors{3.3.1} | FOXQ{3.3.1.17} | 1298228 | 15220 | FOXQ1_MOUSE | O70220 | ||||
| FUBP1_MOUSE.H11MO.0.D |  |
MOUSE:Fubp1 | 12 | D | 0 | TYGTnhKTbKKK | HOCOMOCOv9 | Integrative | 102 | 1196294 | 51886 | FUBP1_MOUSE | Q91WJ8 | ||||||
| GABPA_MOUSE.H11MO.0.A |  |
MOUSE:Gabpa | 14 | A | 0 | vvvvSCGGAAGYRv | HOCOMOCOv11 | ChIP-Seq | 0.988 | 0.952 | 92 | 8 | 500 | Ets-related factors{3.5.2} | Ets-like factors{3.5.2.1} | 95610 | 14390 | GABPA_MOUSE | Q00422 |
| GATA1_MOUSE.H11MO.0.A |  |
MOUSE:Gata1 | 19 | A | 0 | bKbnnvnndvWGATAASvv | HOCOMOCOv11 | ChIP-Seq | 0.964 | 0.951 | 48 | 122 | 500 | GATA-type zinc fingers{2.2.1} | Two zinc-finger GATA factors{2.2.1.1} | 95661 | 14460 | GATA1_MOUSE | P17679 |
| GATA1_MOUSE.H11MO.1.A |  |
MOUSE:Gata1 | 11 | A | 1 | dbAGATAASvv | HOCOMOCOv11 | ChIP-Seq | 0.972 | 0.95 | 48 | 122 | 500 | GATA-type zinc fingers{2.2.1} | Two zinc-finger GATA factors{2.2.1.1} | 95661 | 14460 | GATA1_MOUSE | P17679 |
| GATA2_MOUSE.H11MO.0.A |  |
MOUSE:Gata2 | 11 | A | 0 | dvAGATAASvd | HOCOMOCOv11 | ChIP-Seq | 0.946 | 0.963 | 84 | 54 | 500 | GATA-type zinc fingers{2.2.1} | Two zinc-finger GATA factors{2.2.1.1} | 95662 | 14461 | GATA2_MOUSE | O09100 |
| GATA3_MOUSE.H11MO.0.A |  |
MOUSE:Gata3 | 13 | A | 0 | dvAGATARvvdbh | HOCOMOCOv11 | ChIP-Seq | 0.903 | 0.912 | 98 | 60 | 500 | GATA-type zinc fingers{2.2.1} | Two zinc-finger GATA factors{2.2.1.1} | 95663 | 14462 | GATA3_MOUSE | P23772 |
| GATA4_MOUSE.H11MO.0.A |  |
MOUSE:Gata4 | 10 | A | 0 | dvWGATAASR | HOCOMOCOv11 | ChIP-Seq | 0.955 | 0.94 | 15 | 17 | 500 | GATA-type zinc fingers{2.2.1} | Two zinc-finger GATA factors{2.2.1.1} | 95664 | 14463 | GATA4_MOUSE | Q08369 |
| GATA5_MOUSE.H11MO.0.D |  |
MOUSE:Gata5 | 17 | D | 0 | WvMnWGATAWbnhdRhd | HOCOMOCOv9 | Integrative | 13 | GATA-type zinc fingers{2.2.1} | Two zinc-finger GATA factors{2.2.1.1} | 109497 | 14464 | GATA5_MOUSE | P97489 | ||||
| GATA6_MOUSE.H11MO.0.A |  |
MOUSE:Gata6 | 10 | A | 0 | dvWGATAASv | HOCOMOCOv11 | ChIP-Seq | 0.958 | 0.968 | 29 | 7 | 500 | GATA-type zinc fingers{2.2.1} | Two zinc-finger GATA factors{2.2.1.1} | 107516 | 14465 | GATA6_MOUSE | Q61169 |
| GCM1_MOUSE.H11MO.0.D |  |
MOUSE:Gcm1 | 14 | D | 0 | hWnATGMKGGYhbb | HOCOMOCOv9 | Integrative | 12 | GCM factors{7.2.1} | GCMa (GCM1){7.2.1.0.1} | 108045 | 14531 | GCM1_MOUSE | P70348 | ||||
| GCR_MOUSE.H11MO.0.A |  |
MOUSE:Nr3c1 | 15 | A | 0 | RGdACAbAvWGThCY | HOCOMOCOv11 | ChIP-Seq | 0.978 | 0.99 | 143 | 119 | 500 | Steroid hormone receptors (NR3){2.1.1} | GR-like receptors (NR3C){2.1.1.1} | 95824 | GCR_MOUSE | P06537 | |
| GCR_MOUSE.H11MO.1.A |  |
MOUSE:Nr3c1 | 12 | A | 1 | hvRGRACAbdvh | HOCOMOCOv11 | ChIP-Seq | 0.93 | 0.925 | 143 | 119 | 500 | Steroid hormone receptors (NR3){2.1.1} | GR-like receptors (NR3C){2.1.1.1} | 95824 | GCR_MOUSE | P06537 | |
| GFI1B_MOUSE.H11MO.0.A |  |
MOUSE:Gfi1b | 10 | A | 0 | dYWSTGRTTK | HOCOMOCOv11 | ChIP-Seq | 0.862 | 0.888 | 3 | 24 | 503 | More than 3 adjacent zinc finger factors{2.3.3} | GFI1 factors{2.3.3.21} | 1276578 | 14582 | GFI1B_MOUSE | O70237 |
| GFI1_MOUSE.H11MO.0.C |  |
MOUSE:Gfi1 | 10 | C | 0 | RShSWGATTb | HOCOMOCOv9 | Integrative | 152 | More than 3 adjacent zinc finger factors{2.3.3} | GFI1 factors{2.3.3.21} | 103170 | 14581 | GFI1_MOUSE | P70338 | ||||
| GLI1_MOUSE.H11MO.0.C |  |
MOUSE:Gli1 | 12 | C | 0 | bSTGGGTGGYCh | HOCOMOCOv11 | ChIP-Seq | 0.561 | 0.974 | 2 | 3 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | GLI-like factors{2.3.3.1} | 95727 | 14632 | GLI1_MOUSE | P47806 |
| GLI2_MOUSE.H11MO.0.D |  |
MOUSE:Gli2 | 11 | D | 0 | STGGGTGGTCb | HOCOMOCOv9 | Integrative | 970 | More than 3 adjacent zinc finger factors{2.3.3} | GLI-like factors{2.3.3.1} | 95728 | 14633 | GLI2_MOUSE | Q0VGT2 | ||||
| GLI3_MOUSE.H11MO.0.D |  |
MOUSE:Gli3 | 12 | D | 0 | bbWGKGTGGbYb | HOCOMOCOv11 | ChIP-Seq | 0.52 | 0.752 | 2 | 3 | 476 | More than 3 adjacent zinc finger factors{2.3.3} | GLI-like factors{2.3.3.1} | 95729 | 14634 | GLI3_MOUSE | Q61602 |
| GLIS3_MOUSE.H11MO.0.D |  |
MOUSE:Glis3 | 10 | D | 0 | SYGGGGGGTM | HOCOMOCOv9 | Integrative | 38 | More than 3 adjacent zinc finger factors{2.3.3} | GLI-like factors{2.3.3.1} | 2444289 | 226075 | GLIS3_MOUSE | Q6XP49 | ||||
| GRHL2_MOUSE.H11MO.0.A |  |
MOUSE:Grhl2 | 12 | A | 0 | RRCYdGTYYddC | HOCOMOCOv11 | ChIP-Seq | 0.951 | 0.951 | 22 | 4 | 500 | Grainyhead-related factors{6.7.1} | GRH-like proteins{6.7.1.1} | 2182543 | 252973 | GRHL2_MOUSE | Q8K5C0 |
| HAND1_MOUSE.H11MO.0.C |  |
MOUSE:Hand1 | 17 | C | 0 | ddbYCTGShhWnYvTCW | HOCOMOCOv11 | ChIP-Seq | 0.995 | 3 | 500 | Tal-related factors{1.2.3} | Twist-like factors{1.2.3.2} | 103577 | 15110 | HAND1_MOUSE | Q64279 | ||
| HAND1_MOUSE.H11MO.1.C |  |
MOUSE:Hand1 | 7 | C | 1 | dSYCTGS | HOCOMOCOv11 | ChIP-Seq | 0.951 | 3 | 500 | Tal-related factors{1.2.3} | Twist-like factors{1.2.3.2} | 103577 | 15110 | HAND1_MOUSE | Q64279 | ||
| HBP1_MOUSE.H11MO.0.D |  |
MOUSE:Hbp1 | 9 | D | 0 | RnYSATTGA | HOCOMOCOv9 | Integrative | 5 | SOX-related factors{4.1.1} | Further Sox-related factors{4.1.1.9} | 894659 | 73389 | HBP1_MOUSE | Q8R316 | ||||
| HEN1_MOUSE.H11MO.0.C |  |
MOUSE:Nhlh1 | 20 | C | 0 | bKGGnMbCAGCTGCGKCYhh | HOCOMOCOv9 | Integrative | 55 | Tal-related factors{1.2.3} | Tal / HEN-like factors{1.2.3.1} | 98481 | 18071 | HEN1_MOUSE | Q02576 | ||||
| HES1_MOUSE.H11MO.0.D |  |
MOUSE:Hes1 | 13 | D | 0 | vMSvCAMKvGMhv | HOCOMOCOv9 | Integrative | 28 | Hairy-related factors{1.2.4} | Hairy-like factors{1.2.4.1} | 104853 | 15205 | HES1_MOUSE | P35428 | ||||
| HESX1_MOUSE.H11MO.0.D |  |
MOUSE:Hesx1 | 16 | D | 0 | nKCYGRCWCnTRRnhY | HOCOMOCOv9 | Integrative | 8 | Paired-related HD factors{3.1.3} | HESX{3.1.3.10} | 96071 | 15209 | HESX1_MOUSE | Q61658 | ||||
| HEY2_MOUSE.H11MO.0.D |  |
MOUSE:Hey2 | 17 | D | 0 | GKKSGCWCGTGSCnYKv | HOCOMOCOv9 | Integrative | 8 | Hairy-related factors{1.2.4} | Hairy-like factors{1.2.4.1} | 1341884 | 15214 | HEY2_MOUSE | Q9QUS4 | ||||
| HIC1_MOUSE.H11MO.0.C |  |
MOUSE:Hic1 | 9 | C | 0 | dKGKTGCCM | HOCOMOCOv9 | Integrative | 40 | Factors with multiple dispersed zinc fingers{2.3.4} | Hypermethylated in Cancer proteins{2.3.4.17} | 1338010 | 15248 | HIC1_MOUSE | Q9R1Y5 | ||||
| HIF1A_MOUSE.H11MO.0.C |  |
MOUSE:Hif1a | 8 | C | 0 | vdACGTGh | HOCOMOCOv11 | ChIP-Seq | 0.762 | 0.738 | 39 | 8 | 501 | PAS domain factors{1.2.5} | Ahr-like factors{1.2.5.1} | 106918 | 15251 | HIF1A_MOUSE | Q61221 |
| HINFP_MOUSE.H11MO.0.C |  |
MOUSE:Hinfp | 16 | C | 0 | dhSnnvdCGGACGTWv | HOCOMOCOv9 | Integrative | 42 | Factors with multiple dispersed zinc fingers{2.3.4} | HINFP-like factors{2.3.4.21} | 2429620 | 102423 | HINFP_MOUSE | Q8K1K9 | ||||
| HLF_MOUSE.H11MO.0.C |  |
MOUSE:Hlf | 11 | C | 0 | dvTTRCAYAAb | HOCOMOCOv11 | ChIP-Seq | 0.97 | 4 | 389 | C/EBP-related{1.1.8} | PAR factors{1.1.8.2} | 96108 | 217082 | HLF_MOUSE | Q8BW74 | ||
| HLTF_MOUSE.H11MO.0.D |  |
MOUSE:Hltf | 12 | D | 0 | nAnGKCWGCAAM | HOCOMOCOv9 | Integrative | 10 | 1196437 | 20585 | HLTF_MOUSE | Q6PCN7 | ||||||
| HMGA1_MOUSE.H11MO.0.D |  |
MOUSE:Hmga1 | 7 | D | 0 | nWATTWd | HOCOMOCOv9 | Integrative | 97 | HMGA factors{8.2.1} | HMGA1 (HMGI(Y)){8.2.1.0.1} | 96160 | 111241; 15361 | HMGA1_MOUSE | P17095 | ||||
| HMGA2_MOUSE.H11MO.0.D |  |
MOUSE:Hmga2 | 15 | D | 0 | AATAWhSSSSAATAT | HOCOMOCOv9 | Integrative | 75 | HMGA factors{8.2.1} | HMGA2 (HMGI-C){8.2.1.0.2} | 101761 | 15364 | HMGA2_MOUSE | P52927 | ||||
| HNF1A_MOUSE.H11MO.0.A |  |
MOUSE:Hnf1a | 15 | A | 0 | RGTTAWTvWTYdhYv | HOCOMOCOv9 | Integrative | 175 | POU domain factors{3.1.10} | HNF1-like factors{3.1.10.7} | 98504 | 21405 | HNF1A_MOUSE | P22361 | ||||
| HNF1B_MOUSE.H11MO.0.A |  |
MOUSE:Hnf1b | 15 | A | 0 | dRTWAATGWKTWMYh | HOCOMOCOv11 | ChIP-Seq | 0.947 | 0.995 | 5 | 8 | 500 | POU domain factors{3.1.10} | HNF1-like factors{3.1.10.7} | 98505 | 21410 | HNF1B_MOUSE | P27889 |
| HNF1B_MOUSE.H11MO.1.A |  |
MOUSE:Hnf1b | 13 | A | 1 | hhdGTTAATvWTT | HOCOMOCOv11 | ChIP-Seq | 0.915 | 0.992 | 5 | 8 | 500 | POU domain factors{3.1.10} | HNF1-like factors{3.1.10.7} | 98505 | 21410 | HNF1B_MOUSE | P27889 |
| HNF4A_MOUSE.H11MO.0.A |  |
MOUSE:Hnf4a | 14 | A | 0 | vdRdbCAAAGTYCR | HOCOMOCOv11 | ChIP-Seq | 0.983 | 0.982 | 49 | 33 | 500 | RXR-related receptors (NR2){2.1.3} | HNF-4 (NR2A){2.1.3.2} | 109128 | 15378 | HNF4A_MOUSE | P49698 |
| HNF4G_MOUSE.H11MO.0.C |  |
MOUSE:Hnf4g | 14 | C | 0 | vRGdnCAAAGKbCR | HOCOMOCOv11 | ChIP-Seq | 0.983 | 6 | 500 | RXR-related receptors (NR2){2.1.3} | HNF-4 (NR2A){2.1.3.2} | 1353604 | 30942 | HNF4G_MOUSE | Q9WUU6 | ||
| HNF6_MOUSE.H11MO.0.A |  |
MOUSE:Onecut1 | 11 | A | 0 | hbYATTGATYd | HOCOMOCOv11 | ChIP-Seq | 0.976 | 24 | 512 | HD-CUT factors{3.1.9} | ONECUT{3.1.9.1} | 1196423 | 15379 | HNF6_MOUSE | O08755 | ||
| HSF1_MOUSE.H11MO.0.A |  |
MOUSE:Hsf1 | 15 | A | 0 | bRGAAnnTTCYRGAA | HOCOMOCOv11 | ChIP-Seq | 0.996 | 0.989 | 56 | 32 | 502 | HSF factors{3.4.1} | HSF1 (HSTF1){3.4.1.0.1} | 96238 | 15499 | HSF1_MOUSE | P38532 |
| HSF1_MOUSE.H11MO.1.A |  |
MOUSE:Hsf1 | 9 | A | 1 | bTTCTRGAA | HOCOMOCOv11 | ChIP-Seq | 0.954 | 0.934 | 56 | 32 | 166 | HSF factors{3.4.1} | HSF1 (HSTF1){3.4.1.0.1} | 96238 | 15499 | HSF1_MOUSE | P38532 |
| HSF2_MOUSE.H11MO.0.A |  |
MOUSE:Hsf2 | 14 | A | 0 | vSRWbvWKSKvGRh | HOCOMOCOv9 | Integrative | 1043 | HSF factors{3.4.1} | HSF2 (HSTF2){3.4.1.0.2} | 96239 | 15500 | HSF2_MOUSE | P38533 | ||||
| HTF4_MOUSE.H11MO.0.A |  |
MOUSE:Tcf12 | 10 | A | 0 | dSCAGSTGKb | HOCOMOCOv11 | ChIP-Seq | 0.897 | 0.986 | 49 | 10 | 501 | E2A-related factors{1.2.1} | HTF-4 (TCF-12, HEB){1.2.1.0.3} | 101877 | 21406 | HTF4_MOUSE | Q61286 |
| HXA10_MOUSE.H11MO.0.C |  |
MOUSE:Hoxa10 | 12 | C | 0 | dnYnRTTTATGh | HOCOMOCOv9 | Integrative | 10 | HOX-related factors{3.1.1} | HOX9-13{3.1.1.8} | 96171 | 15395 | HXA10_MOUSE | P31310 | ||||
| HXA13_MOUSE.H11MO.0.C |  |
MOUSE:Hoxa13 | 11 | C | 0 | vdTWTTATKGS | HOCOMOCOv9 | Integrative | 18 | HOX-related factors{3.1.1} | HOX9-13{3.1.1.8} | 96173 | 15398 | HXA13_MOUSE | Q62424 | ||||
| HXA1_MOUSE.H11MO.0.C |  |
MOUSE:Hoxa1 | 10 | C | 0 | nTGATGGATG | HOCOMOCOv9 | Integrative | 8 | HOX-related factors{3.1.1} | HOX1{3.1.1.1} | 96170 | HXA1_MOUSE | P09022 | |||||
| HXA5_MOUSE.H11MO.0.D |  |
MOUSE:Hoxa5 | 10 | D | 0 | YTvATTARYR | HOCOMOCOv9 | Integrative | 54 | HOX-related factors{3.1.1} | HOX5{3.1.1.5} | 96177 | 15402 | HXA5_MOUSE | P09021 | ||||
| HXA7_MOUSE.H11MO.0.D | 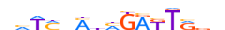 |
MOUSE:Hoxa7 | 15 | D | 0 | vhTMnMnMGAKTRvv | HOCOMOCOv9 | Integrative | 21 | HOX-related factors{3.1.1} | HOX6-7{3.1.1.6} | 96179 | 15404 | HXA7_MOUSE | P02830 | ||||
| HXA9_MOUSE.H11MO.0.B |  |
MOUSE:Hoxa9 | 11 | B | 0 | dTGATTKAYdd | HOCOMOCOv11 | ChIP-Seq | 0.883 | 0.757 | 5 | 8 | 500 | HOX-related factors{3.1.1} | HOX9-13{3.1.1.8} | 96180 | 15405 | HXA9_MOUSE | P09631 |
| HXB1_MOUSE.H11MO.0.D |  |
MOUSE:Hoxb1 | 10 | D | 0 | WGATGGATGG | HOCOMOCOv9 | Integrative | 15 | HOX-related factors{3.1.1} | HOX1{3.1.1.1} | 96182 | 15407 | HXB1_MOUSE | P17919 | ||||
| HXB4_MOUSE.H11MO.0.A |  |
MOUSE:Hoxb4 | 8 | A | 0 | bTAATKRS | HOCOMOCOv11 | ChIP-Seq | 0.491 | 0.894 | 2 | 12 | 502 | HOX-related factors{3.1.1} | HOX4{3.1.1.4} | 96185 | 15412 | HXB4_MOUSE | P10284 |
| HXB6_MOUSE.H11MO.0.D |  |
MOUSE:Hoxb6 | 13 | D | 0 | dWTGATTKATGCn | HOCOMOCOv9 | Integrative | 7 | HOX-related factors{3.1.1} | HOX6-7{3.1.1.6} | 96187 | 15414 | HXB6_MOUSE | P09023 | ||||
| HXB7_MOUSE.H11MO.0.C |  |
MOUSE:Hoxb7 | 10 | C | 0 | WTGATTRATK | HOCOMOCOv9 | Integrative | 28 | HOX-related factors{3.1.1} | HOX6-7{3.1.1.6} | 96188 | 15415 | HXB7_MOUSE | P09024 | ||||
| HXB8_MOUSE.H11MO.0.C |  |
MOUSE:Hoxb8 | 11 | C | 0 | TTGATTRATKv | HOCOMOCOv9 | Integrative | 26 | HOX-related factors{3.1.1} | HOX8{3.1.1.7} | 96189 | 15416 | HXB8_MOUSE | P09632 | ||||
| HXC6_MOUSE.H11MO.0.D |  |
MOUSE:Hoxc6 | 15 | D | 0 | WTGATTTATTWCTTT | HOCOMOCOv9 | Integrative | 7 | HOX-related factors{3.1.1} | HOX6-7{3.1.1.6} | 96197 | 15425 | HXC6_MOUSE | P10629 | ||||
| HXC8_MOUSE.H11MO.0.D |  |
MOUSE:Hoxc8 | 14 | D | 0 | nTTSATTRAnKSnn | HOCOMOCOv9 | Integrative | 9 | HOX-related factors{3.1.1} | HOX8{3.1.1.7} | 96198 | 15426 | HXC8_MOUSE | P09025 | ||||
| HXC9_MOUSE.H11MO.0.A |  |
MOUSE:Hoxc9 | 12 | A | 0 | WRATTdAYdRbh | HOCOMOCOv11 | ChIP-Seq | 0.955 | 7 | 500 | HOX-related factors{3.1.1} | HOX9-13{3.1.1.8} | 96199 | 15427 | HXC9_MOUSE | P09633 | ||
| HXD10_MOUSE.H11MO.0.D |  |
MOUSE:Hoxd10 | 10 | D | 0 | TGnTYTWRTK | HOCOMOCOv9 | Integrative | 7 | HOX-related factors{3.1.1} | HOX9-13{3.1.1.8} | 96202 | 15430 | HXD10_MOUSE | P28359 | ||||
| HXD13_MOUSE.H11MO.0.D |  |
MOUSE:Hoxd13 | 11 | D | 0 | WTTAYKRKRGM | HOCOMOCOv9 | Integrative | 13 | HOX-related factors{3.1.1} | HOX9-13{3.1.1.8} | 96205 | 15433 | HXD13_MOUSE | P70217 | ||||
| HXD4_MOUSE.H11MO.0.D |  |
MOUSE:Hoxd4 | 8 | D | 0 | KTAATTnW | HOCOMOCOv9 | Integrative | 8 | HOX-related factors{3.1.1} | HOX4{3.1.1.4} | 96208 | 15436 | HXD4_MOUSE | P10628 | ||||
| HXD9_MOUSE.H11MO.0.D |  |
MOUSE:Hoxd9 | 10 | D | 0 | AnWTTnATnn | HOCOMOCOv9 | Integrative | 6 | HOX-related factors{3.1.1} | HOX9-13{3.1.1.8} | 96210 | 15438 | HXD9_MOUSE | P28357 | ||||
| IKZF1_MOUSE.H11MO.0.C |  |
MOUSE:Ikzf1 | 8 | C | 0 | bTGGGARd | HOCOMOCOv9 | Integrative | 61 | Factors with multiple dispersed zinc fingers{2.3.4} | Ikaros{2.3.4.4} | 1342540 | IKZF1_MOUSE | Q03267 | |||||
| INSM1_MOUSE.H11MO.0.C |  |
MOUSE:Insm1 | 12 | C | 0 | KGYMAGGGGGCA | HOCOMOCOv9 | Integrative | 24 | Factors with multiple dispersed zinc fingers{2.3.4} | Insulinoma-associated proteins{2.3.4.16} | 1859980 | 53626 | INSM1_MOUSE | Q63ZV0 | ||||
| IRF1_MOUSE.H11MO.0.A | 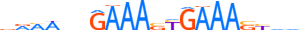 |
MOUSE:Irf1 | 20 | A | 0 | vRMWnhGAAASYGAAASYvd | HOCOMOCOv11 | ChIP-Seq | 0.997 | 0.999 | 23 | 18 | 504 | Interferon-regulatory factors{3.5.3} | IRF-1{3.5.3.0.1} | 96590 | 16362 | IRF1_MOUSE | P15314 |
| IRF2_MOUSE.H11MO.0.B |  |
MOUSE:Irf2 | 20 | B | 0 | nRRWvhGAAASYGAAASYvv | HOCOMOCOv11 | ChIP-Seq | 0.994 | 8 | 499 | Interferon-regulatory factors{3.5.3} | IRF-2{3.5.3.0.2} | 96591 | 16363 | IRF2_MOUSE | P23906 | ||
| IRF3_MOUSE.H11MO.0.A |  |
MOUSE:Irf3 | 20 | A | 0 | dRRRddRAAAvdGRAASnRR | HOCOMOCOv11 | ChIP-Seq | 0.789 | 0.948 | 6 | 21 | 505 | Interferon-regulatory factors{3.5.3} | IRF-3{3.5.3.0.3} | 1859179 | 54131 | IRF3_MOUSE | P70671 |
| IRF4_MOUSE.H11MO.0.A |  |
MOUSE:Irf4 | 18 | A | 0 | ddRWvvRGAAvTGARAvh | HOCOMOCOv11 | ChIP-Seq | 0.951 | 0.981 | 19 | 61 | 500 | Interferon-regulatory factors{3.5.3} | IRF-4 (LSIRF, NF-EM5, MUM1, Pip){3.5.3.0.4} | 1096873 | 16364 | IRF4_MOUSE | Q64287 |
| IRF5_MOUSE.H11MO.0.D |  |
MOUSE:Irf5 | 20 | D | 0 | KAAAGGAAAGCnAAAAGTGM | HOCOMOCOv9 | Integrative | 5 | Interferon-regulatory factors{3.5.3} | IRF-5{3.5.3.0.5} | 1350924 | 27056 | IRF5_MOUSE | P56477 | ||||
| IRF7_MOUSE.H11MO.0.C |  |
MOUSE:Irf7 | 10 | C | 0 | RAAAbYRAAW | HOCOMOCOv9 | Integrative | 40 | Interferon-regulatory factors{3.5.3} | IRF-7{3.5.3.0.7} | 1859212 | 54123 | IRF7_MOUSE | P70434 | ||||
| IRF8_MOUSE.H11MO.0.A |  |
MOUSE:Irf8 | 20 | A | 0 | vdddvRGGAASTGAAASYdv | HOCOMOCOv11 | ChIP-Seq | 0.996 | 59 | 505 | Interferon-regulatory factors{3.5.3} | IRF-8 (ICSBP1){3.5.3.0.8} | 96395 | 15900 | IRF8_MOUSE | P23611 | ||
| IRF9_MOUSE.H11MO.0.C |  |
MOUSE:Irf9 | 12 | C | 0 | GAAAbhGAAACT | HOCOMOCOv9 | Integrative | 5 | Interferon-regulatory factors{3.5.3} | IRF-9 (ISGF-3gamma){3.5.3.0.9} | 107587 | 16391 | IRF9_MOUSE | Q61179 | ||||
| ISL1_MOUSE.H11MO.0.A |  |
MOUSE:Isl1 | 9 | A | 0 | vYWRAKKRv | HOCOMOCOv11 | ChIP-Seq | 0.805 | 0.858 | 3 | 13 | 500 | HD-LIM factors{3.1.5} | ISL{3.1.5.1} | 101791 | 16392 | ISL1_MOUSE | P61372 |
| ITF2_MOUSE.H11MO.0.B |  |
MOUSE:Tcf4 | 9 | B | 0 | vCAGGTGYd | HOCOMOCOv9 | Integrative | 553 | E2A-related factors{1.2.1} | SEF2 (E2-2, TCF-4, ITF-2, MITF){1.2.1.0.2} | 98506 | 21413 | ITF2_MOUSE | Q60722 | ||||
| JUNB_MOUSE.H11MO.0.A |  |
MOUSE:Junb | 9 | A | 0 | nTGAGTCAb | HOCOMOCOv11 | ChIP-Seq | 0.977 | 0.935 | 8 | 25 | 500 | Jun-related factors{1.1.1} | Jun factors{1.1.1.1} | 96647 | 16477 | JUNB_MOUSE | P09450 |
| JUND_MOUSE.H11MO.0.A |  |
MOUSE:Jund | 11 | A | 0 | vdvTGAGTCAb | HOCOMOCOv11 | ChIP-Seq | 0.995 | 0.988 | 64 | 21 | 450 | Jun-related factors{1.1.1} | Jun factors{1.1.1.1} | 96648 | 16478 | JUND_MOUSE | P15066 |
| JUN_MOUSE.H11MO.0.A |  |
MOUSE:Jun | 10 | A | 0 | dvTGAGTCAb | HOCOMOCOv11 | ChIP-Seq | 0.949 | 0.938 | 56 | 39 | 500 | Jun-related factors{1.1.1} | Jun factors{1.1.1.1} | 96646 | 16476 | JUN_MOUSE | P05627 |
| KAISO_MOUSE.H11MO.0.B |  |
MOUSE:Zbtb33 | 15 | B | 0 | vdRvbMYCGCGAGAv | HOCOMOCOv11 | ChIP-Seq | 0.9 | 37 | 498 | Other factors with up to three adjacent zinc fingers{2.3.2} | Factors with 2-3 adjacent zinc fingersand a BTB/POZ domain{2.3.2.1} | 1927290 | 56805 | KAISO_MOUSE | Q8BN78 | ||
| KAISO_MOUSE.H11MO.1.B |  |
MOUSE:Zbtb33 | 11 | B | 1 | hvYCGCGAGAn | HOCOMOCOv11 | ChIP-Seq | 0.888 | 37 | 502 | Other factors with up to three adjacent zinc fingers{2.3.2} | Factors with 2-3 adjacent zinc fingersand a BTB/POZ domain{2.3.2.1} | 1927290 | 56805 | KAISO_MOUSE | Q8BN78 | ||
| KAISO_MOUSE.H11MO.2.B |  |
MOUSE:Zbtb33 | 12 | B | 2 | hTvGCARGAddn | HOCOMOCOv9 | Integrative | 0.841 | 37 | 2038 | Other factors with up to three adjacent zinc fingers{2.3.2} | Factors with 2-3 adjacent zinc fingersand a BTB/POZ domain{2.3.2.1} | 1927290 | 56805 | KAISO_MOUSE | Q8BN78 | ||
| KLF15_MOUSE.H11MO.0.A |  |
MOUSE:Klf15 | 19 | A | 0 | RRGGMGGRGvnRGGRRdSR | HOCOMOCOv11 | ChIP-Seq | 0.952 | 0.998 | 4 | 3 | 481 | Three-zinc finger Krüppel-related factors{2.3.1} | Krüppel-like factors{2.3.1.2} | 1929988 | 66277 | KLF15_MOUSE | Q9EPW2 |
| KLF1_MOUSE.H11MO.0.A |  |
MOUSE:Klf1 | 14 | A | 0 | dGGGYGKGGShbSK | HOCOMOCOv11 | ChIP-Seq | 0.893 | 0.982 | 5 | 6 | 454 | Three-zinc finger Krüppel-related factors{2.3.1} | Krüppel-like factors{2.3.1.2} | 1342771 | KLF1_MOUSE | P46099 | |
| KLF3_MOUSE.H11MO.0.A |  |
MOUSE:Klf3 | 19 | A | 0 | vvvvvdGGGYGGRGbhbbv | HOCOMOCOv11 | ChIP-Seq | 0.76 | 0.992 | 2 | 7 | 498 | Three-zinc finger Krüppel-related factors{2.3.1} | Krüppel-like factors{2.3.1.2} | 1342773 | 16599 | KLF3_MOUSE | Q60980 |
| KLF4_MOUSE.H11MO.0.A |  |
MOUSE:Klf4 | 15 | A | 0 | dvvdvWGGGYGKGKM | HOCOMOCOv11 | ChIP-Seq | 0.909 | 0.975 | 11 | 34 | 500 | Three-zinc finger Krüppel-related factors{2.3.1} | Krüppel-like factors{2.3.1.2} | 1342287 | 16600 | KLF4_MOUSE | Q60793 |
| KLF5_MOUSE.H11MO.0.A |  |
MOUSE:Klf5 | 15 | A | 0 | dRvvvdGGGYGKGGM | HOCOMOCOv11 | ChIP-Seq | 0.943 | 0.985 | 32 | 23 | 500 | Three-zinc finger Krüppel-related factors{2.3.1} | Krüppel-like factors{2.3.1.2} | 1338056 | 12224 | KLF5_MOUSE | Q9Z0Z7 |
| KLF6_MOUSE.H11MO.0.B |  |
MOUSE:Klf6 | 19 | B | 0 | vvvRGGGYGKRGvnbRSvv | HOCOMOCOv11 | ChIP-Seq | 0.946 | 0.604 | 16 | 2 | 498 | Three-zinc finger Krüppel-related factors{2.3.1} | Krüppel-like factors{2.3.1.2} | 1346318 | KLF6_MOUSE | O08584 | |
| KLF8_MOUSE.H11MO.0.C |  |
MOUSE:Klf8 | 9 | C | 0 | CAGGGGGTG | HOCOMOCOv9 | Integrative | 5 | Three-zinc finger Krüppel-related factors{2.3.1} | Krüppel-like factors{2.3.1.2} | 2442430 | 245671 | KLF8_MOUSE | Q8BLM0 | ||||
| LEF1_MOUSE.H11MO.0.B |  |
MOUSE:Lef1 | 14 | B | 0 | bSYTTTSWhnKbhh | HOCOMOCOv11 | ChIP-Seq | 0.876 | 20 | 456 | TCF-7-related factors{4.1.3} | LEF-1 (TCF-1alpha) [1]{4.1.3.0.4} | 96770 | 16842 | LEF1_MOUSE | P27782 | ||
| LHX2_MOUSE.H11MO.0.A |  |
MOUSE:Lhx2 | 10 | A | 0 | nhTAATKAGn | HOCOMOCOv11 | ChIP-Seq | 0.814 | 0.938 | 5 | 12 | 501 | HD-LIM factors{3.1.5} | Lhx-2-like factors{3.1.5.3} | 96785 | 16870 | LHX2_MOUSE | Q9Z0S2 |
| LHX3_MOUSE.H11MO.0.C |  |
MOUSE:Lhx3 | 9 | C | 0 | nbTAATKRv | HOCOMOCOv11 | ChIP-Seq | 0.94 | 2 | 500 | HD-LIM factors{3.1.5} | Lhx-3-like factors{3.1.5.4} | 102673 | 16871 | LHX3_MOUSE | P50481 | ||
| LHX6_MOUSE.H11MO.0.C |  |
MOUSE:Lhx6 | 9 | C | 0 | nYTGATTRv | HOCOMOCOv11 | ChIP-Seq | 0.517 | 0.946 | 1 | 2 | 241 | HD-LIM factors{3.1.5} | Lhx-6-like factors{3.1.5.5} | 1306803 | 16874 | LHX6_MOUSE | Q9R1R0 |
| LYL1_MOUSE.H11MO.0.A |  |
MOUSE:Lyl1 | 14 | A | 0 | vMWbMTGYbhYbbb | HOCOMOCOv11 | ChIP-Seq | 0.852 | 0.808 | 23 | 3 | 453 | Tal-related factors{1.2.3} | Tal / HEN-like factors{1.2.3.1} | 96891 | 17095 | LYL1_MOUSE | P27792 |
| MAFA_MOUSE.H11MO.0.D |  |
MOUSE:Mafa | 13 | D | 0 | bTGCTGMbdbdbb | HOCOMOCOv11 | ChIP-Seq | 0.799 | 3 | 500 | Maf-related factors{1.1.3} | Large Maf factors{1.1.3.1} | 2673307 | 378435 | MAFA_MOUSE | Q8CF90 | ||
| MAFB_MOUSE.H11MO.0.B |  |
MOUSE:Mafb | 11 | B | 0 | TGCTGAbhhdd | HOCOMOCOv11 | ChIP-Seq | 0.705 | 0.953 | 5 | 3 | 474 | Maf-related factors{1.1.3} | Large Maf factors{1.1.3.1} | 104555 | 16658 | MAFB_MOUSE | P54841 |
| MAFF_MOUSE.H11MO.0.A |  |
MOUSE:Maff | 22 | A | 0 | dRRWnWGCTGASYMRGCddddd | HOCOMOCOv11 | ChIP-Seq | 0.451 | 0.957 | 2 | 15 | 500 | Maf-related factors{1.1.3} | Small Maf factors{1.1.3.2} | 96910 | 17133 | MAFF_MOUSE | O54791 |
| MAFF_MOUSE.H11MO.1.A |  |
MOUSE:Maff | 10 | A | 1 | TGCTGASTMA | HOCOMOCOv11 | ChIP-Seq | 0.503 | 0.929 | 2 | 15 | 500 | Maf-related factors{1.1.3} | Small Maf factors{1.1.3.2} | 96910 | 17133 | MAFF_MOUSE | O54791 |
| MAFG_MOUSE.H11MO.0.A |  |
MOUSE:Mafg | 18 | A | 0 | ddWWnTGCTGASTCAGCW | HOCOMOCOv11 | ChIP-Seq | 0.869 | 0.998 | 3 | 6 | 430 | Maf-related factors{1.1.3} | Small Maf factors{1.1.3.2} | 96911 | 17134 | MAFG_MOUSE | O54790 |
| MAFG_MOUSE.H11MO.1.A |  |
MOUSE:Mafg | 8 | A | 1 | TGCTGWSK | HOCOMOCOv11 | ChIP-Seq | 0.874 | 0.989 | 3 | 6 | 500 | Maf-related factors{1.1.3} | Small Maf factors{1.1.3.2} | 96911 | 17134 | MAFG_MOUSE | O54790 |
| MAFK_MOUSE.H11MO.0.A |  |
MOUSE:Mafk | 20 | A | 0 | ddWhbTGCTGASTCAKSdhd | HOCOMOCOv11 | ChIP-Seq | 0.977 | 0.988 | 12 | 11 | 441 | Maf-related factors{1.1.3} | Small Maf factors{1.1.3.2} | 99951 | 17135 | MAFK_MOUSE | Q61827 |
| MAFK_MOUSE.H11MO.1.A |  |
MOUSE:Mafk | 13 | A | 1 | WGCTGAGTCAKSW | HOCOMOCOv11 | ChIP-Seq | 0.958 | 0.979 | 12 | 11 | 390 | Maf-related factors{1.1.3} | Small Maf factors{1.1.3.2} | 99951 | 17135 | MAFK_MOUSE | Q61827 |
| MAF_MOUSE.H11MO.0.A |  |
MOUSE:Maf | 17 | A | 0 | dhdbTGCTGAbdhhKbh | HOCOMOCOv11 | ChIP-Seq | 0.846 | 0.853 | 10 | 5 | 488 | Maf-related factors{1.1.3} | Large Maf factors{1.1.3.1} | 96909 | 17132 | MAF_MOUSE | P54843 |
| MAF_MOUSE.H11MO.1.A |  |
MOUSE:Maf | 9 | A | 1 | nTGCTGASh | HOCOMOCOv11 | ChIP-Seq | 0.765 | 0.834 | 10 | 5 | 500 | Maf-related factors{1.1.3} | Large Maf factors{1.1.3.1} | 96909 | 17132 | MAF_MOUSE | P54843 |
| MAX_MOUSE.H11MO.0.A |  |
MOUSE:Max | 11 | A | 0 | vvvCACGYGSv | HOCOMOCOv11 | ChIP-Seq | 0.939 | 0.961 | 86 | 14 | 505 | bHLH-ZIP factors{1.2.6} | Myc / Max factors{1.2.6.5} | 96921 | MAX_MOUSE | P28574 | |
| MAZ_MOUSE.H11MO.0.A |  |
MOUSE:Maz | 22 | A | 0 | vRRGGWGGRRSnRRRRRvdvdR | HOCOMOCOv11 | ChIP-Seq | 0.951 | 0.961 | 3 | 5 | 501 | Factors with multiple dispersed zinc fingers{2.3.4} | MAZ-like factors{2.3.4.8} | 1338823 | MAZ_MOUSE | P56671 | |
| MAZ_MOUSE.H11MO.1.A |  |
MOUSE:Maz | 11 | A | 1 | RRGGGMGGGGv | HOCOMOCOv11 | ChIP-Seq | 0.926 | 0.955 | 3 | 5 | 486 | Factors with multiple dispersed zinc fingers{2.3.4} | MAZ-like factors{2.3.4.8} | 1338823 | MAZ_MOUSE | P56671 | |
| MBD2_MOUSE.H11MO.0.B |  |
MOUSE:Mbd2 | 11 | B | 0 | nSGGCCGGMKv | HOCOMOCOv9 | Integrative | 113 | 1333813 | 17191 | MBD2_MOUSE | Q9Z2E1 | ||||||
| MCR_MOUSE.H11MO.0.D |  |
MOUSE:Nr3c2 | 17 | D | 0 | hAGWMCARSTKGTdGKR | HOCOMOCOv9 | Integrative | 8 | Steroid hormone receptors (NR3){2.1.1} | GR-like receptors (NR3C){2.1.1.1} | 99459 | MCR_MOUSE | Q8VII8 | |||||
| MECP2_MOUSE.H11MO.0.C |  |
MOUSE:Mecp2 | 7 | C | 0 | SCCGGAG | HOCOMOCOv9 | Integrative | 122 | 99918 | 17257 | MECP2_MOUSE | Q9Z2D6 | ||||||
| MEF2A_MOUSE.H11MO.0.A |  |
MOUSE:Mef2a | 14 | A | 0 | ddYTATTTWWRKhh | HOCOMOCOv11 | ChIP-Seq | 0.94 | 0.984 | 15 | 6 | 498 | Regulators of differentiation{5.1.1} | MEF-2{5.1.1.1} | 99532 | 17258 | MEF2A_MOUSE | Q60929 |
| MEF2C_MOUSE.H11MO.0.A |  |
MOUSE:Mef2c | 15 | A | 0 | dddYTATTTWWRKhh | HOCOMOCOv11 | ChIP-Seq | 0.956 | 0.929 | 6 | 9 | 264 | Regulators of differentiation{5.1.1} | MEF-2{5.1.1.1} | 99458 | 17260 | MEF2C_MOUSE | Q8CFN5 |
| MEF2D_MOUSE.H11MO.0.A |  |
MOUSE:Mef2d | 14 | A | 0 | dKYTATTTWTAKhh | HOCOMOCOv11 | ChIP-Seq | 0.812 | 0.97 | 2 | 27 | 500 | Regulators of differentiation{5.1.1} | MEF-2{5.1.1.1} | 99533 | 17261 | MEF2D_MOUSE | Q63943 |
| MEIS1_MOUSE.H11MO.0.A |  |
MOUSE:Meis1 | 12 | A | 0 | dTGATWdATddb | HOCOMOCOv11 | ChIP-Seq | 0.816 | 0.945 | 2 | 37 | 499 | TALE-type homeo domain factors{3.1.4} | MEIS{3.1.4.2} | 104717 | 17268 | MEIS1_MOUSE | Q60954 |
| MEIS1_MOUSE.H11MO.1.A |  |
MOUSE:Meis1 | 12 | A | 1 | bWShTGACAKvh | HOCOMOCOv11 | ChIP-Seq | 0.708 | 0.891 | 2 | 37 | 500 | TALE-type homeo domain factors{3.1.4} | MEIS{3.1.4.2} | 104717 | 17268 | MEIS1_MOUSE | Q60954 |
| MEIS2_MOUSE.H11MO.0.B |  |
MOUSE:Meis2 | 13 | B | 0 | hTGACAGbTKTbv | HOCOMOCOv9 | Integrative | 413 | TALE-type homeo domain factors{3.1.4} | MEIS{3.1.4.2} | 108564 | 17536 | MEIS2_MOUSE | P97367 | ||||
| MITF_MOUSE.H11MO.0.A |  |
MOUSE:Mitf | 10 | A | 0 | bChYGTGMYh | HOCOMOCOv11 | ChIP-Seq | 0.996 | 0.835 | 28 | 3 | 501 | bHLH-ZIP factors{1.2.6} | TFE3-like factors{1.2.6.1} | 104554 | 17342 | MITF_MOUSE | Q08874 |
| MLXPL_MOUSE.H11MO.0.D |  |
MOUSE:Mlxipl | 19 | D | 0 | SCAYGKGRMMMYnbSGTGb | HOCOMOCOv9 | Integrative | 11 | bHLH-ZIP factors{1.2.6} | Mondo-like factors{1.2.6.6} | 1927999 | 58805 | MLXPL_MOUSE | Q99MZ3 | ||||
| MSGN1_MOUSE.H11MO.0.C |  |
MOUSE:Msgn1 | 12 | C | 0 | nCCATTTGKYhh | HOCOMOCOv11 | ChIP-Seq | 0.999 | 4 | 500 | Tal-related factors{1.2.3} | Mesp-like factors{1.2.3.3} | 1860483 | 56184 | MSGN1_MOUSE | Q9JK54 | ||
| MSX2_MOUSE.H11MO.0.D |  |
MOUSE:Msx2 | 7 | D | 0 | TAATTnb | HOCOMOCOv9 | Integrative | 12 | NK-related factors{3.1.2} | MSX{3.1.2.11} | 97169 | 17702 | MSX2_MOUSE | Q03358 | ||||
| MSX3_MOUSE.H11MO.0.D |  |
MOUSE:Msx3 | 17 | D | 0 | TAATTGbWWdWTAATTd | HOCOMOCOv10 | HT-SELEX | 2675 | 106587 | 17703 | MSX3_MOUSE | P70354 | ||||||
| MTF1_MOUSE.H11MO.0.C |  |
MOUSE:Mtf1 | 17 | C | 0 | vGKRMCGKGTGCAdMdb | HOCOMOCOv9 | Integrative | 33 | More than 3 adjacent zinc finger factors{2.3.3} | MTF1-like factors{2.3.3.24} | 101786 | 17764 | MTF1_MOUSE | Q07243 | ||||
| MXI1_MOUSE.H11MO.0.A | 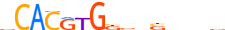 |
MOUSE:Mxi1 | 15 | A | 0 | vCACRYGKvnSbbbv | HOCOMOCOv10 | ChIP-Seq | 0.878 | 0.973 | 10 | 5 | 500 | bHLH-ZIP factors{1.2.6} | Mad-like factors{1.2.6.7} | 97245 | 17859 | MXI1_MOUSE | P50540 |
| MXI1_MOUSE.H11MO.1.A |  |
MOUSE:Mxi1 | 8 | A | 1 | hCRCGTGS | HOCOMOCOv11 | ChIP-Seq | 0.856 | 0.973 | 10 | 5 | 510 | bHLH-ZIP factors{1.2.6} | Mad-like factors{1.2.6.7} | 97245 | 17859 | MXI1_MOUSE | P50540 |
| MYBA_MOUSE.H11MO.0.C |  |
MOUSE:Mybl1 | 15 | C | 0 | vbbGRCvGTTRvnvv | HOCOMOCOv11 | ChIP-Seq | 0.533 | 0.897 | 3 | 2 | 500 | Myb/SANT domain factors{3.5.1} | Myb-like factors{3.5.1.1} | 99925 | 17864 | MYBA_MOUSE | P51960 |
| MYBB_MOUSE.H11MO.0.D |  |
MOUSE:Mybl2 | 10 | D | 0 | dRSAnGTTdW | HOCOMOCOv9 | Integrative | 9 | Myb/SANT domain factors{3.5.1} | Myb-like factors{3.5.1.1} | 101785 | 17865 | MYBB_MOUSE | P48972 | ||||
| MYB_MOUSE.H11MO.0.A |  |
MOUSE:Myb | 12 | A | 0 | dvdKRCvGTTRv | HOCOMOCOv11 | ChIP-Seq | 0.83 | 0.934 | 27 | 9 | 500 | Myb/SANT domain factors{3.5.1} | Myb-like factors{3.5.1.1} | 97249 | 17863 | MYB_MOUSE | P06876 |
| MYCN_MOUSE.H11MO.0.A |  |
MOUSE:Mycn | 8 | A | 0 | SCRCGTGK | HOCOMOCOv11 | ChIP-Seq | 0.897 | 0.928 | 32 | 10 | 508 | bHLH-ZIP factors{1.2.6} | Myc / Max factors{1.2.6.5} | 97357 | 18109 | MYCN_MOUSE | P03966 |
| MYC_MOUSE.H11MO.0.A |  |
MOUSE:Myc | 12 | A | 0 | vvMCWCGTGSbS | HOCOMOCOv11 | ChIP-Seq | 0.917 | 0.957 | 265 | 126 | 501 | bHLH-ZIP factors{1.2.6} | Myc / Max factors{1.2.6.5} | 97250 | 17869 | MYC_MOUSE | P01108 |
| MYF5_MOUSE.H11MO.0.D |  |
MOUSE:Myf5 | 11 | D | 0 | bCASCTGTYnh | HOCOMOCOv11 | ChIP-Seq | 0.787 | 2 | 449 | MyoD / ASC-related factors{1.2.2} | Myogenic transcription factors{1.2.2.1} | 97252 | 17877 | MYF5_MOUSE | P24699 | ||
| MYF6_MOUSE.H11MO.0.C |  |
MOUSE:Myf6 | 7 | C | 0 | CAGCTGC | HOCOMOCOv9 | Integrative | 9 | MyoD / ASC-related factors{1.2.2} | Myogenic transcription factors{1.2.2.1} | 97253 | 17878 | MYF6_MOUSE | P15375 | ||||
| MYOD1_MOUSE.H11MO.0.A |  |
MOUSE:Myod1 | 15 | A | 0 | vCASCTGYYnYhvbb | HOCOMOCOv11 | ChIP-Seq | 0.985 | 0.994 | 9 | 63 | 501 | MyoD / ASC-related factors{1.2.2} | Myogenic transcription factors{1.2.2.1} | 97275 | 17927 | MYOD1_MOUSE | P10085 |
| MYOD1_MOUSE.H11MO.1.A |  |
MOUSE:Myod1 | 11 | A | 1 | vCAGCTGTYvY | HOCOMOCOv11 | ChIP-Seq | 0.982 | 0.992 | 9 | 63 | 501 | MyoD / ASC-related factors{1.2.2} | Myogenic transcription factors{1.2.2.1} | 97275 | 17927 | MYOD1_MOUSE | P10085 |
| MYOG_MOUSE.H11MO.0.A |  |
MOUSE:Myog | 13 | A | 0 | bhvCAGCTGYYhb | HOCOMOCOv11 | ChIP-Seq | 0.994 | 20 | 500 | MyoD / ASC-related factors{1.2.2} | Myogenic transcription factors{1.2.2.1} | 97276 | 17928 | MYOG_MOUSE | P12979 | ||
| NANOG_MOUSE.H11MO.0.A |  |
MOUSE:Nanog | 17 | A | 0 | bbYWTTSWnWTKYWRAd | HOCOMOCOv11 | ChIP-Seq | 0.914 | 0.939 | 15 | 38 | 500 | NK-related factors{3.1.2} | NANOG{3.1.2.12} | 1919200 | 71950 | NANOG_MOUSE | Q80Z64 |
| NANOG_MOUSE.H11MO.1.A |  |
MOUSE:Nanog | 11 | A | 1 | vRSCCATYAvn | HOCOMOCOv11 | ChIP-Seq | 0.769 | 0.825 | 15 | 38 | 500 | NK-related factors{3.1.2} | NANOG{3.1.2.12} | 1919200 | 71950 | NANOG_MOUSE | Q80Z64 |
| NDF1_MOUSE.H11MO.0.A |  |
MOUSE:Neurod1 | 10 | A | 0 | RRCAGMTGGb | HOCOMOCOv11 | ChIP-Seq | 0.946 | 0.952 | 5 | 2 | 503 | Tal-related factors{1.2.3} | Neurogenin / Atonal-like factors{1.2.3.4} | 1339708 | 18012 | NDF1_MOUSE | Q60867 |
| NDF2_MOUSE.H11MO.0.A |  |
MOUSE:Neurod2 | 10 | A | 0 | RRCAGATGGb | HOCOMOCOv11 | ChIP-Seq | 0.543 | 0.961 | 2 | 21 | 503 | Tal-related factors{1.2.3} | Neurogenin / Atonal-like factors{1.2.3.4} | 107755 | 18013 | NDF2_MOUSE | Q62414 |
| NF2L1_MOUSE.H11MO.0.C |  |
MOUSE:Nfe2l1 | 7 | C | 0 | hRTCAYn | HOCOMOCOv9 | Integrative | 115 | Jun-related factors{1.1.1} | NF-E2-like factors{1.1.1.2} | 99421 | 18023 | NF2L1_MOUSE | Q61985 | ||||
| NF2L2_MOUSE.H11MO.0.A |  |
MOUSE:Nfe2l2 | 14 | A | 0 | RTGASWhRGCWddh | HOCOMOCOv11 | ChIP-Seq | 0.894 | 0.986 | 5 | 26 | 478 | Jun-related factors{1.1.1} | NF-E2-like factors{1.1.1.2} | 108420 | 18024 | NF2L2_MOUSE | Q60795 |
| NFAC1_MOUSE.H11MO.0.A |  |
MOUSE:Nfatc1 | 15 | A | 0 | WGGAAAndnTKWvhM | HOCOMOCOv11 | ChIP-Seq | 0.714 | 0.955 | 7 | 8 | 445 | NFAT-related factors{6.1.3} | NFATc1{6.1.3.0.1} | 102469 | NFAC1_MOUSE | O88942 | |
| NFAC1_MOUSE.H11MO.1.A |  |
MOUSE:Nfatc1 | 11 | A | 1 | vvvWGGAAAvn | HOCOMOCOv11 | ChIP-Seq | 0.689 | 0.908 | 7 | 8 | 500 | NFAT-related factors{6.1.3} | NFATc1{6.1.3.0.1} | 102469 | NFAC1_MOUSE | O88942 | |
| NFAC2_MOUSE.H11MO.0.C |  |
MOUSE:Nfatc2 | 9 | C | 0 | dGGAAAWhb | HOCOMOCOv9 | Integrative | 0.915 | 4 | 87 | NFAT-related factors{6.1.3} | NFATc2 (NFATp, NFAT1){6.1.3.0.2} | 102463 | 18019 | NFAC2_MOUSE | Q60591 | ||
| NFAC2_MOUSE.H11MO.1.C |  |
MOUSE:Nfatc2 | 7 | C | 1 | vWGGAAR | HOCOMOCOv11 | ChIP-Seq | 0.934 | 4 | 499 | NFAT-related factors{6.1.3} | NFATc2 (NFATp, NFAT1){6.1.3.0.2} | 102463 | 18019 | NFAC2_MOUSE | Q60591 | ||
| NFAC3_MOUSE.H11MO.0.B |  |
MOUSE:Nfatc3 | 9 | B | 0 | hGGAAAAhY | HOCOMOCOv9 | Integrative | 18 | NFAT-related factors{6.1.3} | NFATc3 (NFATx, NFAT4){6.1.3.0.3} | 103296 | NFAC3_MOUSE | P97305 | |||||
| NFAC4_MOUSE.H11MO.0.C |  |
MOUSE:Nfatc4 | 10 | C | 0 | dGGAAARhdW | HOCOMOCOv9 | Integrative | 19 | NFAT-related factors{6.1.3} | NFATc4 (NFAT3){6.1.3.0.4} | 1920431 | 73181 | NFAC4_MOUSE | Q8K120 | ||||
| NFAT5_MOUSE.H11MO.0.D |  |
MOUSE:Nfat5 | 15 | D | 0 | STGGAAAAnYYCMKK | HOCOMOCOv9 | Integrative | 8 | NFAT-related factors{6.1.3} | NFAT5 (TonEBP){6.1.3.0.5} | 1859333 | 54446 | NFAT5_MOUSE | Q9WV30 | ||||
| NFE2_MOUSE.H11MO.0.A |  |
MOUSE:Nfe2 | 14 | A | 0 | RTGASTCAGCAddh | HOCOMOCOv11 | ChIP-Seq | 0.905 | 0.992 | 17 | 9 | 500 | Jun-related factors{1.1.1} | NF-E2-like factors{1.1.1.2} | 97308 | 18022 | NFE2_MOUSE | Q07279 |
| NFIA_MOUSE.H11MO.0.C |  |
MOUSE:Nfia | 15 | C | 0 | hTGGShnGnhKCCWd | HOCOMOCOv9 | Integrative | 233 | Nuclear factor 1{7.1.2} | NF-1A (NF-IA){7.1.2.0.1} | 108056 | 18027 | NFIA_MOUSE | Q02780 | ||||
| NFIA_MOUSE.H11MO.1.D |  |
MOUSE:Nfia | 7 | D | 1 | hYTGGCd | HOCOMOCOv9 | Integrative | 267 | Nuclear factor 1{7.1.2} | NF-1A (NF-IA){7.1.2.0.1} | 108056 | 18027 | NFIA_MOUSE | Q02780 | ||||
| NFIB_MOUSE.H11MO.0.C |  |
MOUSE:Nfib | 9 | C | 0 | YYTGGCWbv | HOCOMOCOv11 | ChIP-Seq | 0.596 | 0.832 | 2 | 2 | 500 | Nuclear factor 1{7.1.2} | NF-1B{7.1.2.0.2} | 103188 | 18028 | NFIB_MOUSE | P97863 |
| NFIC_MOUSE.H11MO.0.A |  |
MOUSE:Nfic | 9 | A | 0 | bbTTGGCWb | HOCOMOCOv11 | ChIP-Seq | 0.932 | 0.831 | 18 | 1 | 500 | Nuclear factor 1{7.1.2} | NF-1C{7.1.2.0.3} | 109591 | 18029 | NFIC_MOUSE | P70255 |
| NFIC_MOUSE.H11MO.1.A |  |
MOUSE:Nfic | 15 | A | 1 | bYTGGCWbnnnbhYh | HOCOMOCOv11 | ChIP-Seq | 0.958 | 0.807 | 18 | 1 | 410 | Nuclear factor 1{7.1.2} | NF-1C{7.1.2.0.3} | 109591 | 18029 | NFIC_MOUSE | P70255 |
| NFIL3_MOUSE.H11MO.0.C |  |
MOUSE:Nfil3 | 11 | C | 0 | dvTTATGYAAb | HOCOMOCOv11 | ChIP-Seq | 0.543 | 0.869 | 2 | 2 | 501 | C/EBP-related{1.1.8} | PAR factors{1.1.8.2} | 109495 | 18030 | NFIL3_MOUSE | O08750 |
| NFKB1_MOUSE.H11MO.0.A |  |
MOUSE:Nfkb1 | 11 | A | 0 | GGGAAAbYCCC | HOCOMOCOv11 | ChIP-Seq | 0.726 | 0.953 | 7 | 16 | 503 | NF-kappaB-related factors{6.1.1} | NF-kappaB p50 subunit-like factors{6.1.1.1} | 97312 | 18033 | NFKB1_MOUSE | P25799 |
| NFKB2_MOUSE.H11MO.0.C |  |
MOUSE:Nfkb2 | 11 | C | 0 | KGGRAAdYCCC | HOCOMOCOv11 | ChIP-Seq | 0.977 | 4 | 500 | NF-kappaB-related factors{6.1.1} | NF-kappaB p50 subunit-like factors{6.1.1.1} | 1099800 | 18034 | NFKB2_MOUSE | Q9WTK5 | ||
| NFYA_MOUSE.H11MO.0.A |  |
MOUSE:Nfya | 14 | A | 0 | bbRRCCAATSRvMR | HOCOMOCOv11 | ChIP-Seq | 0.982 | 0.96 | 19 | 14 | 500 | Heteromeric CCAAT-binding factors{4.2.1} | NF-YA (CP1A, CBF-B){4.2.1.0.1} | 97316 | 18044 | NFYA_MOUSE | P23708 |
| NFYB_MOUSE.H11MO.0.A |  |
MOUSE:Nfyb | 13 | A | 0 | bRRCCAATSRSvd | HOCOMOCOv9 | Integrative | 0.987 | 0.997 | 4 | 4 | 1163 | Heteromeric CCAAT-binding factors{4.2.1} | NF-YB (CP1B, CBF-A){4.2.1.0.2} | 97317 | 18045 | NFYB_MOUSE | P63139 |
| NFYC_MOUSE.H11MO.0.B |  |
MOUSE:Nfyc | 14 | B | 0 | bbRRCCAATSRRMR | HOCOMOCOv11 | ChIP-Seq | 0.695 | 0.994 | 4 | 4 | 500 | Heteromeric CCAAT-binding factors{4.2.1} | NF-YC{4.2.1.0.3} | 107901 | 18046 | NFYC_MOUSE | P70353 |
| NGN2_MOUSE.H11MO.0.C | 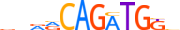 |
MOUSE:Neurog2 | 12 | C | 0 | vbRRCAGATGKb | HOCOMOCOv11 | ChIP-Seq | 0.977 | 4 | 206 | Tal-related factors{1.2.3} | Neurogenin / Atonal-like factors{1.2.3.4} | 109619 | 11924 | NGN2_MOUSE | P70447 | ||
| NKX21_MOUSE.H11MO.0.A |  |
MOUSE:Nkx2-1 | 10 | A | 0 | bbWKGAGWGb | HOCOMOCOv11 | ChIP-Seq | 0.923 | 0.92 | 14 | 9 | 500 | NK-related factors{3.1.2} | NK-2.1{3.1.2.14} | 108067 | 21869 | NKX21_MOUSE | P50220 |
| NKX22_MOUSE.H11MO.0.A |  |
MOUSE:Nkx2-2 | 11 | A | 0 | bTbRAGWGbbb | HOCOMOCOv11 | ChIP-Seq | 0.399 | 0.82 | 2 | 5 | 500 | NK-related factors{3.1.2} | NK-2.2{3.1.2.15} | 97347 | 18088 | NKX22_MOUSE | P42586 |
| NKX25_MOUSE.H11MO.0.A |  |
MOUSE:Nkx2-5 | 10 | A | 0 | nbYbRAGWGS | HOCOMOCOv11 | ChIP-Seq | 0.612 | 0.896 | 2 | 7 | 500 | NK-related factors{3.1.2} | NK-4{3.1.2.17} | 97350 | 18091 | NKX25_MOUSE | P42582 |
| NKX28_MOUSE.H11MO.0.C |  |
MOUSE:Nkx2-8 | 9 | C | 0 | bTCAAGGAb | HOCOMOCOv9 | Integrative | 6 | NK-related factors{3.1.2} | NK-2.2{3.1.2.15} | 1270158 | 18094 | NKX28_MOUSE | O70584 | ||||
| NKX31_MOUSE.H11MO.0.C |  |
MOUSE:Nkx3-1 | 12 | C | 0 | nbdRAGTGbYWd | HOCOMOCOv11 | ChIP-Seq | 0.792 | 0.662 | 11 | 10 | 501 | NK-related factors{3.1.2} | NK-3{3.1.2.16} | 97352 | 18095 | NKX31_MOUSE | P97436 |
| NKX32_MOUSE.H11MO.0.B |  |
MOUSE:Nkx3-2 | 10 | B | 0 | WTAAGTGbhh | HOCOMOCOv11 | ChIP-Seq | 0.982 | 6 | 501 | NK-related factors{3.1.2} | NK-3{3.1.2.16} | 108015 | 12020 | NKX32_MOUSE | P97503 | ||
| NKX61_MOUSE.H11MO.0.A |  |
MOUSE:Nkx6-1 | 19 | A | 0 | MhWdATKdvhWWWWRAWdd | HOCOMOCOv11 | ChIP-Seq | 0.379 | 0.953 | 3 | 8 | 498 | NK-related factors{3.1.2} | NK-6{3.1.2.19} | 1206039 | 18096 | NKX61_MOUSE | Q99MA9 |
| NKX61_MOUSE.H11MO.1.A |  |
MOUSE:Nkx6-1 | 9 | A | 1 | dWhAATGRv | HOCOMOCOv11 | ChIP-Seq | 0.448 | 0.867 | 3 | 8 | 500 | NK-related factors{3.1.2} | NK-6{3.1.2.19} | 1206039 | 18096 | NKX61_MOUSE | Q99MA9 |
| NOBOX_MOUSE.H11MO.0.C |  |
MOUSE:Nobox | 9 | C | 0 | hTAATTGvb | HOCOMOCOv9 | Integrative | 66 | 108011 | 18291 | NOBOX_MOUSE | Q8VIH1 | ||||||
| NR0B1_MOUSE.H11MO.0.D |  |
MOUSE:Nr0b1 | 10 | D | 0 | SCGKGGRASA | HOCOMOCOv9 | Integrative | 9 | DAX-related receptors (NR0){2.1.7} | DAX1 (NR0B1){2.1.7.0.1} | 1352460 | 11614 | NR0B1_MOUSE | Q61066 | ||||
| NR1D1_MOUSE.H11MO.0.A |  |
MOUSE:Nr1d1 | 19 | A | 0 | dRvnMWbWGKGdMAvdddv | HOCOMOCOv11 | ChIP-Seq | 0.615 | 0.883 | 1 | 53 | 501 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Rev-ErbA (NR1D){2.1.2.3} | 2444210 | 217166 | NR1D1_MOUSE | Q3UV55 |
| NR1D1_MOUSE.H11MO.1.C |  |
MOUSE:Nr1d1 | 14 | C | 1 | WWAAvTAGGTCAnd | HOCOMOCOv9 | Integrative | 0.419 | 0.759 | 1 | 53 | 13 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Rev-ErbA (NR1D){2.1.2.3} | 2444210 | 217166 | NR1D1_MOUSE | Q3UV55 |
| NR1D2_MOUSE.H11MO.0.A |  |
MOUSE:Nr1d2 | 19 | A | 0 | dvRdMWvKRRGKCAvdvdd | HOCOMOCOv11 | ChIP-Seq | 0.892 | 12 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Rev-ErbA (NR1D){2.1.2.3} | 2449205 | 353187 | NR1D2_MOUSE | Q60674 | ||
| NR1H2_MOUSE.H11MO.0.D |  |
MOUSE:Nr1h2 | 19 | D | 0 | TMRAGGTCAAAGGTCAASK | HOCOMOCOv9 | Integrative | 24 | Thyroid hormone receptor-related factors (NR1){2.1.2} | LXR (NR1H){2.1.2.7} | 1352463 | 22260 | NR1H2_MOUSE | Q60644 | ||||
| NR1H3_MOUSE.H11MO.0.A |  |
MOUSE:Nr1h3 | 15 | A | 0 | vdRdnMMRAGKYCAb | HOCOMOCOv11 | ChIP-Seq | 0.606 | 0.896 | 13 | 10 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | LXR (NR1H){2.1.2.7} | 1352462 | 22259 | NR1H3_MOUSE | Q9Z0Y9 |
| NR1H3_MOUSE.H11MO.1.A |  |
MOUSE:Nr1h3 | 19 | A | 1 | vhvRGKYhAbhnnRGKdYA | HOCOMOCOv11 | ChIP-Seq | 0.77 | 0.86 | 13 | 10 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | LXR (NR1H){2.1.2.7} | 1352462 | 22259 | NR1H3_MOUSE | Q9Z0Y9 |
| NR1H3_MOUSE.H11MO.2.A |  |
MOUSE:Nr1h3 | 10 | A | 2 | hvAGGKCAvd | HOCOMOCOv11 | ChIP-Seq | 0.761 | 0.903 | 13 | 10 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | LXR (NR1H){2.1.2.7} | 1352462 | 22259 | NR1H3_MOUSE | Q9Z0Y9 |
| NR1H4_MOUSE.H11MO.0.A |  |
MOUSE:Nr1h4 | 19 | A | 0 | vRGKKSRvYGdCCbbbvGK | HOCOMOCOv11 | ChIP-Seq | 0.926 | 15 | 451 | Thyroid hormone receptor-related factors (NR1){2.1.2} | LXR (NR1H){2.1.2.7} | 1352464 | 20186 | NR1H4_MOUSE | Q60641 | ||
| NR1H4_MOUSE.H11MO.1.A |  |
MOUSE:Nr1h4 | 8 | A | 1 | RGGKCAbh | HOCOMOCOv11 | ChIP-Seq | 0.849 | 15 | 501 | Thyroid hormone receptor-related factors (NR1){2.1.2} | LXR (NR1H){2.1.2.7} | 1352464 | 20186 | NR1H4_MOUSE | Q60641 | ||
| NR1I2_MOUSE.H11MO.0.C |  |
MOUSE:Nr1i2 | 19 | C | 0 | ddRRSTYARnnRAGKTCAb | HOCOMOCOv9 | Integrative | 29 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Vitamin D receptor (NR1I){2.1.2.4} | 1337040 | 18171 | NR1I2_MOUSE | O54915 | ||||
| NR1I2_MOUSE.H11MO.1.D |  |
MOUSE:Nr1i2 | 8 | D | 1 | dAGKTCAb | HOCOMOCOv9 | Integrative | 45 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Vitamin D receptor (NR1I){2.1.2.4} | 1337040 | 18171 | NR1I2_MOUSE | O54915 | ||||
| NR1I3_MOUSE.H11MO.0.C |  |
MOUSE:Nr1i3 | 18 | C | 0 | RRGSTCAvvvRAGKWYRd | HOCOMOCOv9 | Integrative | 28 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Vitamin D receptor (NR1I){2.1.2.4} | 1346307 | 12355 | NR1I3_MOUSE | O35627 | ||||
| NR1I3_MOUSE.H11MO.1.D |  |
MOUSE:Nr1i3 | 8 | D | 1 | RAGTTCAb | HOCOMOCOv9 | Integrative | 33 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Vitamin D receptor (NR1I){2.1.2.4} | 1346307 | 12355 | NR1I3_MOUSE | O35627 | ||||
| NR2C1_MOUSE.H11MO.0.C |  |
MOUSE:Nr2c1 | 13 | C | 0 | RbYvARAGGTCAn | HOCOMOCOv9 | Integrative | 19 | RXR-related receptors (NR2){2.1.3} | Testicular receptors (NR2C){2.1.3.4} | 1352465 | 22025 | NR2C1_MOUSE | Q505F1 | ||||
| NR2C2_MOUSE.H11MO.0.A |  |
MOUSE:Nr2c2 | 13 | A | 0 | RRSbSARAGGKMR | HOCOMOCOv9 | Integrative | 2062 | RXR-related receptors (NR2){2.1.3} | Testicular receptors (NR2C){2.1.3.4} | 1352466 | 22026 | NR2C2_MOUSE | P49117 | ||||
| NR2E3_MOUSE.H11MO.0.C |  |
MOUSE:Nr2e3 | 14 | C | 0 | nAAKTCAAAARKMM | HOCOMOCOv9 | Integrative | 23 | RXR-related receptors (NR2){2.1.3} | Tailless-like receptors (NR2E){2.1.3.3} | 1346317 | 23958 | NR2E3_MOUSE | Q9QXZ7 | ||||
| NR2F6_MOUSE.H11MO.0.D |  |
MOUSE:Nr2f6 | 18 | D | 0 | RGSWMAAAGdnSAnnKdR | HOCOMOCOv9 | Integrative | 11 | RXR-related receptors (NR2){2.1.3} | COUP-like receptors (NR2F){2.1.3.5} | 1352453 | 13864 | NR2F6_MOUSE | P43136 | ||||
| NR4A1_MOUSE.H11MO.0.B |  |
MOUSE:Nr4a1 | 9 | B | 0 | dRARdGbMA | HOCOMOCOv11 | ChIP-Seq | 0.903 | 0.631 | 12 | 2 | 499 | NGFI-B-related receptors (NR4){2.1.4} | NGFI-B (Nur77, NR4A1){2.1.4.0.1} | 1352454 | 15370 | NR4A1_MOUSE | P12813 |
| NR4A2_MOUSE.H11MO.0.C |  |
MOUSE:Nr4a2 | 9 | C | 0 | RAAGGTCAv | HOCOMOCOv9 | Integrative | 33 | NGFI-B-related receptors (NR4){2.1.4} | NURR1 (NR4A2){2.1.4.0.2} | 1352456 | 18227 | NR4A2_MOUSE | Q06219 | ||||
| NR4A3_MOUSE.H11MO.0.D |  |
MOUSE:Nr4a3 | 10 | D | 0 | CAAAGGTCAS | HOCOMOCOv9 | Integrative | 5 | NGFI-B-related receptors (NR4){2.1.4} | NOR1 (NR4A3, TEC){2.1.4.0.3} | 1352457 | 18124 | NR4A3_MOUSE | Q9QZB6 | ||||
| NR5A2_MOUSE.H11MO.0.A |  |
MOUSE:Nr5a2 | 11 | A | 0 | dbYCAAGGYCR | HOCOMOCOv11 | ChIP-Seq | 0.537 | 0.957 | 4 | 8 | 500 | FTZ-F1-related receptors (NR5){2.1.5} | LRH-1 (NR5A2){2.1.5.0.2} | 1346834 | 26424 | NR5A2_MOUSE | P45448 |
| NR6A1_MOUSE.H11MO.0.D |  |
MOUSE:Nr6a1 | 13 | D | 0 | dRKKYCAAGKbYW | HOCOMOCOv11 | ChIP-Seq | 0.784 | 5 | 501 | GCNF-related receptors (NR6){2.1.6} | GCNF (NR6A1){2.1.6.0.1} | 1352459 | 14536 | NR6A1_MOUSE | Q64249 | ||
| NRF1_MOUSE.H11MO.0.A |  |
MOUSE:Nrf1 | 17 | A | 0 | YdvYGCGCMKGCGCvvv | HOCOMOCOv11 | ChIP-Seq | 0.999 | 0.997 | 31 | 23 | 500 | NRF{0.0.6} | NRF-1 (alpha-pal){0.0.6.0.1} | 1332235 | 18181 | NRF1_MOUSE | Q9WU00 |
| OLIG2_MOUSE.H11MO.0.A |  |
MOUSE:Olig2 | 18 | A | 0 | MCAKCTGYYbbbhdbhhb | HOCOMOCOv11 | ChIP-Seq | 0.925 | 19 | 500 | Tal-related factors{1.2.3} | Neurogenin / Atonal-like factors{1.2.3.4} | 1355331 | 50913 | OLIG2_MOUSE | Q9EQW6 | ||
| OLIG2_MOUSE.H11MO.1.A |  |
MOUSE:Olig2 | 10 | A | 1 | hMWKMTGYYb | HOCOMOCOv11 | ChIP-Seq | 0.905 | 19 | 500 | Tal-related factors{1.2.3} | Neurogenin / Atonal-like factors{1.2.3.4} | 1355331 | 50913 | OLIG2_MOUSE | Q9EQW6 | ||
| ONEC2_MOUSE.H11MO.0.D |  |
MOUSE:Onecut2 | 21 | D | 0 | RKvhYdTbATKGATTWTWTTW | HOCOMOCOv9 | Integrative | 7 | HD-CUT factors{3.1.9} | ONECUT{3.1.9.1} | 1891408 | 225631 | ONEC2_MOUSE | Q6XBJ3 | ||||
| OTX1_MOUSE.H11MO.0.D |  |
MOUSE:Otx1 | 8 | D | 0 | nGGATTAn | HOCOMOCOv9 | Integrative | 8 | Paired-related HD factors{3.1.3} | OTX{3.1.3.17} | 97450 | 18423 | OTX1_MOUSE | P80205 | ||||
| OTX2_MOUSE.H11MO.0.A |  |
MOUSE:Otx2 | 12 | A | 0 | dvddRGGATTAd | HOCOMOCOv11 | ChIP-Seq | 0.847 | 0.904 | 5 | 18 | 500 | Paired-related HD factors{3.1.3} | OTX{3.1.3.17} | 97451 | OTX2_MOUSE | P80206 | |
| OVOL1_MOUSE.H11MO.0.C |  |
MOUSE:Ovol1 | 9 | C | 0 | KGTAACKGT | HOCOMOCOv9 | Integrative | 7 | More than 3 adjacent zinc finger factors{2.3.3} | OVOL-factors{2.3.3.17} | 1330290 | 18426 | OVOL1_MOUSE | Q9WTJ2 | ||||
| OVOL2_MOUSE.H11MO.0.C |  |
MOUSE:Ovol2 | 8 | C | 0 | dTAACGGd | HOCOMOCOv11 | ChIP-Seq | 0.453 | 0.952 | 2 | 2 | 197 | More than 3 adjacent zinc finger factors{2.3.3} | OVOL-factors{2.3.3.17} | 1338039 | 107586 | OVOL2_MOUSE | Q8CIV7 |
| P53_MOUSE.H11MO.0.A |  |
MOUSE:Tp53 | 20 | A | 0 | RRRCAWGYMYRGRCWdRhMh | HOCOMOCOv11 | ChIP-Seq | 0.995 | 0.988 | 141 | 39 | 498 | p53-related factors{6.3.1} | p53{6.3.1.0.1} | 98834 | 22059 | P53_MOUSE | P02340 |
| P53_MOUSE.H11MO.1.A |  |
MOUSE:Tp53 | 13 | A | 1 | dRRCAWGYCYRGR | HOCOMOCOv11 | ChIP-Seq | 0.987 | 0.981 | 141 | 39 | 504 | p53-related factors{6.3.1} | p53{6.3.1.0.1} | 98834 | 22059 | P53_MOUSE | P02340 |
| P63_MOUSE.H11MO.0.B |  |
MOUSE:Tp63 | 19 | B | 0 | dRCRWGYhCWRRCWKGYhb | HOCOMOCOv11 | ChIP-Seq | 0.996 | 79 | 500 | p53-related factors{6.3.1} | p63{6.3.1.0.2} | 1330810 | 22061 | P63_MOUSE | O88898 | ||
| P63_MOUSE.H11MO.1.B |  |
MOUSE:Tp63 | 11 | B | 1 | vdRCMWGYCWR | HOCOMOCOv11 | ChIP-Seq | 0.936 | 79 | 502 | p53-related factors{6.3.1} | p63{6.3.1.0.2} | 1330810 | 22061 | P63_MOUSE | O88898 | ||
| P73_MOUSE.H11MO.0.B |  |
MOUSE:Tp73 | 19 | B | 0 | dddCWWGYYMMdRCWdGYh | HOCOMOCOv11 | ChIP-Seq | 0.996 | 10 | 500 | p53-related factors{6.3.1} | p73{6.3.1.0.3} | 1336991 | 22062 | P73_MOUSE | Q9JJP2 | ||
| P73_MOUSE.H11MO.1.B |  |
MOUSE:Tp73 | 12 | B | 1 | vdRCWWGYCYRR | HOCOMOCOv11 | ChIP-Seq | 0.908 | 10 | 500 | p53-related factors{6.3.1} | p73{6.3.1.0.3} | 1336991 | 22062 | P73_MOUSE | Q9JJP2 | ||
| PAX2_MOUSE.H11MO.0.D |  |
MOUSE:Pax2 | 18 | D | 0 | vhbnvdTSAhRvdYGAYW | HOCOMOCOv9 | Integrative | 50 | Paired domain only{3.2.2} | PAX-2-like factors (partial homeobox){3.2.2.2} | 97486 | 18504 | PAX2_MOUSE | P32114 | ||||
| PAX3_MOUSE.H11MO.0.D |  |
MOUSE:Pax3 | 13 | D | 0 | ndnWMATKGvWKh | HOCOMOCOv11 | ChIP-Seq | 0.773 | 3 | 455 | Paired plus homeo domain{3.2.1} | PAX-3/7{3.2.1.1} | 97487 | 18505 | PAX3_MOUSE | P24610 | ||
| PAX5_MOUSE.H11MO.0.A |  |
MOUSE:Pax5 | 19 | A | 0 | vdRnKYRvYvRdGCRdRRM | HOCOMOCOv11 | ChIP-Seq | 0.944 | 0.861 | 32 | 8 | 500 | Paired domain only{3.2.2} | PAX-2-like factors (partial homeobox){3.2.2.2} | 97489 | 18507 | PAX5_MOUSE | Q02650 |
| PAX6_MOUSE.H11MO.0.C | 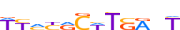 |
MOUSE:Pax6 | 12 | C | 0 | WYMYRCWTSRnT | HOCOMOCOv11 | ChIP-Seq | 0.749 | 0.734 | 6 | 8 | 500 | Paired plus homeo domain{3.2.1} | PAX-4/6{3.2.1.2} | 97490 | 18508 | PAX6_MOUSE | P63015 |
| PAX8_MOUSE.H11MO.0.D |  |
MOUSE:Pax8 | 14 | D | 0 | bYvAYYSRvSYRKd | HOCOMOCOv9 | Integrative | 37 | Paired domain only{3.2.2} | PAX-2-like factors (partial homeobox){3.2.2.2} | 97492 | 18510 | PAX8_MOUSE | Q00288 | ||||
| PBX1_MOUSE.H11MO.0.A |  |
MOUSE:Pbx1 | 11 | A | 0 | WGAKTGRCRKv | HOCOMOCOv11 | ChIP-Seq | 0.858 | 0.902 | 2 | 36 | 500 | TALE-type homeo domain factors{3.1.4} | PBX{3.1.4.4} | 97495 | 18514 | PBX1_MOUSE | P41778 |
| PBX1_MOUSE.H11MO.1.A |  |
MOUSE:Pbx1 | 13 | A | 1 | bbbYSATTGGYhd | HOCOMOCOv11 | ChIP-Seq | 0.876 | 0.915 | 2 | 36 | 500 | TALE-type homeo domain factors{3.1.4} | PBX{3.1.4.4} | 97495 | 18514 | PBX1_MOUSE | P41778 |
| PBX1_MOUSE.H11MO.2.C |  |
MOUSE:Pbx1 | 15 | C | 2 | ddhWWGATKGATbdb | HOCOMOCOv9 | Integrative | 0.733 | 0.759 | 2 | 36 | 145 | TALE-type homeo domain factors{3.1.4} | PBX{3.1.4.4} | 97495 | 18514 | PBX1_MOUSE | P41778 |
| PBX2_MOUSE.H11MO.0.C |  |
MOUSE:Pbx2 | 15 | C | 0 | KvMhKTGATTGWWKb | HOCOMOCOv9 | Integrative | 19 | TALE-type homeo domain factors{3.1.4} | PBX{3.1.4.4} | 1341793 | 18515 | PBX2_MOUSE | O35984 | ||||
| PBX3_MOUSE.H11MO.0.A |  |
MOUSE:Pbx3 | 14 | A | 0 | bbbYSATTGGYddv | HOCOMOCOv9 | Integrative | 1885 | TALE-type homeo domain factors{3.1.4} | PBX{3.1.4.4} | 97496 | 18516 | PBX3_MOUSE | O35317 | ||||
| PDX1_MOUSE.H11MO.0.B |  |
MOUSE:Pdx1 | 9 | B | 0 | vYTAATGRb | HOCOMOCOv11 | ChIP-Seq | 0.752 | 0.895 | 10 | 4 | 500 | HOX-related factors{3.1.1} | PDX{3.1.1.15} | 102851 | 18609 | PDX1_MOUSE | P52946 |
| PEBB_MOUSE.H11MO.0.C |  |
MOUSE:Cbfb | 11 | C | 0 | YbTGTGGTYdb | HOCOMOCOv9 | Integrative | 23 | 99851 | 12400 | PEBB_MOUSE | Q08024 | ||||||
| PIT1_MOUSE.H11MO.0.C |  |
MOUSE:Pou1f1 | 13 | C | 0 | bWMWTGvAWATWh | HOCOMOCOv9 | Integrative | 39 | POU domain factors{3.1.10} | POU1 (Pit-1-like factors){3.1.10.1} | 97588 | 18736 | PIT1_MOUSE | Q00286 | ||||
| PITX1_MOUSE.H11MO.0.C |  |
MOUSE:Pitx1 | 10 | C | 0 | dRGGATTAdv | HOCOMOCOv11 | ChIP-Seq | 0.529 | 0.875 | 2 | 4 | 196 | Paired-related HD factors{3.1.3} | PITX{3.1.3.19} | 107374 | 18740 | PITX1_MOUSE | P70314 |
| PITX2_MOUSE.H11MO.0.D |  |
MOUSE:Pitx2 | 10 | D | 0 | YKGGATTWvh | HOCOMOCOv9 | Integrative | 17 | Paired-related HD factors{3.1.3} | PITX{3.1.3.19} | 109340 | 18741 | PITX2_MOUSE | P97474 | ||||
| PKNX1_MOUSE.H11MO.0.A |  |
MOUSE:Pknox1 | 11 | A | 0 | WGAbKGRCRSv | HOCOMOCOv11 | ChIP-Seq | 0.967 | 19 | 480 | TALE-type homeo domain factors{3.1.4} | PKNOX{3.1.4.5} | 1201409 | 18771 | PKNX1_MOUSE | O70477 | ||
| PLAG1_MOUSE.H11MO.0.D |  |
MOUSE:Plag1 | 17 | D | 0 | RGGGGSMhndRdMGGGK | HOCOMOCOv9 | Integrative | 25 | More than 3 adjacent zinc finger factors{2.3.3} | PLAG factors{2.3.3.25} | 1891916 | 56711 | PLAG1_MOUSE | Q9QYE0 | ||||
| PO2F1_MOUSE.H11MO.0.C |  |
MOUSE:Pou2f1 | 16 | C | 0 | RnKbvhATGSAAWhRh | HOCOMOCOv11 | ChIP-Seq | 0.766 | 0.669 | 15 | 14 | 134 | POU domain factors{3.1.10} | POU2 (Oct-1/2-like factors){3.1.10.2} | 101898 | 18986 | PO2F1_MOUSE | P25425 |
| PO2F2_MOUSE.H11MO.0.B |  |
MOUSE:Pou2f2 | 11 | B | 0 | dhATGCAWAKd | HOCOMOCOv11 | ChIP-Seq | 0.919 | 14 | 501 | POU domain factors{3.1.10} | POU2 (Oct-1/2-like factors){3.1.10.2} | 101897 | 18987 | PO2F2_MOUSE | Q00196 | ||
| PO3F1_MOUSE.H11MO.0.C |  |
MOUSE:Pou3f1 | 16 | C | 0 | bWhATKMAWAWKbMdd | HOCOMOCOv11 | ChIP-Seq | 0.789 | 10 | 500 | POU domain factors{3.1.10} | POU3 (Oct-6-like factors){3.1.10.3} | 101896 | 18991 | PO3F1_MOUSE | P21952 | ||
| PO3F2_MOUSE.H11MO.0.A |  |
MOUSE:Pou3f2 | 16 | A | 0 | hhYATGMATAWKYAWd | HOCOMOCOv11 | ChIP-Seq | 0.788 | 0.943 | 12 | 22 | 500 | POU domain factors{3.1.10} | POU3 (Oct-6-like factors){3.1.10.3} | 101895 | 18992 | PO3F2_MOUSE | P31360 |
| PO3F2_MOUSE.H11MO.1.A |  |
MOUSE:Pou3f2 | 17 | A | 1 | dbSWYMTGAATAWKbAW | HOCOMOCOv11 | ChIP-Seq | 0.777 | 0.934 | 12 | 22 | 284 | POU domain factors{3.1.10} | POU3 (Oct-6-like factors){3.1.10.3} | 101895 | 18992 | PO3F2_MOUSE | P31360 |
| PO4F2_MOUSE.H11MO.0.D |  |
MOUSE:Pou4f2 | 13 | D | 0 | YvTTAATGAKhnn | HOCOMOCOv9 | Integrative | 31 | POU domain factors{3.1.10} | POU4 (Brn-3-like factors){3.1.10.4} | 102524 | 18997 | PO4F2_MOUSE | Q63934 | ||||
| PO5F1_MOUSE.H11MO.0.A | 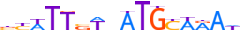 |
MOUSE:Pou5f1 | 16 | A | 0 | bYWTTSWhATGYWRAW | HOCOMOCOv11 | ChIP-Seq | 0.98 | 0.985 | 44 | 119 | 499 | POU domain factors{3.1.10} | POU5 (Oct-3/4-like factors){3.1.10.5} | 101893 | 18999 | PO5F1_MOUSE | P20263 |
| PO5F1_MOUSE.H11MO.1.A |  |
MOUSE:Pou5f1 | 9 | A | 1 | hWTGCWRAh | HOCOMOCOv11 | ChIP-Seq | 0.873 | 0.877 | 44 | 119 | 498 | POU domain factors{3.1.10} | POU5 (Oct-3/4-like factors){3.1.10.5} | 101893 | 18999 | PO5F1_MOUSE | P20263 |
| PO6F1_MOUSE.H11MO.0.D |  |
MOUSE:Pou6f1 | 13 | D | 0 | nATAATTTATGCR | HOCOMOCOv9 | Integrative | 14 | POU domain factors{3.1.10} | POU6 (Brn-5-like factors){3.1.10.6} | 102935 | 19009 | PO6F1_MOUSE | Q07916 | ||||
| PPARA_MOUSE.H11MO.0.A |  |
MOUSE:Ppara | 17 | A | 0 | hWbWRdGbvARAGGKbR | HOCOMOCOv11 | ChIP-Seq | 0.585 | 0.958 | 5 | 20 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | PPAR (NR1C){2.1.2.5} | 104740 | 19013 | PPARA_MOUSE | P23204 |
| PPARA_MOUSE.H11MO.1.A |  |
MOUSE:Ppara | 9 | A | 1 | SARAGGKSA | HOCOMOCOv11 | ChIP-Seq | 0.483 | 0.863 | 5 | 20 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | PPAR (NR1C){2.1.2.5} | 104740 | 19013 | PPARA_MOUSE | P23204 |
| PPARD_MOUSE.H11MO.0.D |  |
MOUSE:Ppard | 14 | D | 0 | YAGKnnAMAGGWYA | HOCOMOCOv9 | Integrative | 18 | Thyroid hormone receptor-related factors (NR1){2.1.2} | PPAR (NR1C){2.1.2.5} | 101884 | 19015 | PPARD_MOUSE | P35396 | ||||
| PPARG_MOUSE.H11MO.0.A |  |
MOUSE:Pparg | 17 | A | 0 | hWbhRKGbSARAGGKhR | HOCOMOCOv11 | ChIP-Seq | 0.959 | 0.96 | 53 | 114 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | PPAR (NR1C){2.1.2.5} | 97747 | 19016 | PPARG_MOUSE | P37238 |
| PPARG_MOUSE.H11MO.1.A |  |
MOUSE:Pparg | 10 | A | 1 | bSARAGGKSA | HOCOMOCOv11 | ChIP-Seq | 0.809 | 0.838 | 53 | 114 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | PPAR (NR1C){2.1.2.5} | 97747 | 19016 | PPARG_MOUSE | P37238 |
| PRD14_MOUSE.H11MO.0.A |  |
MOUSE:Prdm14 | 16 | A | 0 | vddSKTAGRGMMYhdb | HOCOMOCOv11 | ChIP-Seq | 0.858 | 0.928 | 7 | 11 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | unclassified{2.3.3.0} | 3588194 | 383491 | PRD14_MOUSE | E9Q3T6 |
| PRD16_MOUSE.H11MO.0.B |  |
MOUSE:Prdm16 | 10 | B | 0 | bCCCAGRGvv | HOCOMOCOv11 | ChIP-Seq | 0.878 | 3 | 501 | Factors with multiple dispersed zinc fingers{2.3.4} | Evi-1-like factors{2.3.4.14} | 1917923 | 70673 | PRD16_MOUSE | A2A935 | ||
| PRDM1_MOUSE.H11MO.0.A |  |
MOUSE:Prdm1 | 14 | A | 0 | RRRAGdGAAAGKdv | HOCOMOCOv11 | ChIP-Seq | 0.959 | 0.962 | 12 | 9 | 502 | More than 3 adjacent zinc finger factors{2.3.3} | PRDM1-like factors{2.3.3.12} | 99655 | 12142 | PRDM1_MOUSE | Q60636 |
| PRDM5_MOUSE.H11MO.0.A |  |
MOUSE:Prdm5 | 15 | A | 0 | vvdGGWGvdSvdGKh | HOCOMOCOv11 | ChIP-Seq | 0.952 | 12 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | unclassified{2.3.3.0} | 1918029 | 70779 | PRDM5_MOUSE | Q9CXE0 | ||
| PRDM5_MOUSE.H11MO.1.A |  |
MOUSE:Prdm5 | 9 | A | 1 | vMdGGAGvn | HOCOMOCOv11 | ChIP-Seq | 0.855 | 12 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | unclassified{2.3.3.0} | 1918029 | 70779 | PRDM5_MOUSE | Q9CXE0 | ||
| PRDM9_MOUSE.H11MO.0.C |  |
MOUSE:Prdm9 | 19 | C | 0 | hRnddGGYAGYAvCWbCWb | HOCOMOCOv11 | ChIP-Seq | 0.919 | 4 | 490 | Factors with multiple dispersed zinc fingers{2.3.4} | unclassified{2.3.4.0} | 2384854 | PRDM9_MOUSE | Q96EQ9 | |||
| PRGR_MOUSE.H11MO.0.A |  |
MOUSE:Pgr | 15 | A | 0 | RGdACAvAvWGThCY | HOCOMOCOv11 | ChIP-Seq | 0.962 | 0.975 | 51 | 15 | 500 | Steroid hormone receptors (NR3){2.1.1} | GR-like receptors (NR3C){2.1.1.1} | 97567 | PRGR_MOUSE | Q00175 | |
| PRGR_MOUSE.H11MO.1.A |  |
MOUSE:Pgr | 8 | A | 1 | vRGdACAb | HOCOMOCOv11 | ChIP-Seq | 0.845 | 0.924 | 51 | 15 | 500 | Steroid hormone receptors (NR3){2.1.1} | GR-like receptors (NR3C){2.1.1.1} | 97567 | PRGR_MOUSE | Q00175 | |
| PROP1_MOUSE.H11MO.0.C |  |
MOUSE:Prop1 | 11 | C | 0 | WAATKGvRTTW | HOCOMOCOv11 | ChIP-Seq | 0.921 | 3 | 500 | Paired-related HD factors{3.1.3} | PROP{3.1.3.20} | 109330 | 19127 | PROP1_MOUSE | P97458 | ||
| PRRX1_MOUSE.H11MO.0.D |  |
MOUSE:Prrx1 | 7 | D | 0 | TAATCTG | HOCOMOCOv9 | Integrative | 4 | Paired-related HD factors{3.1.3} | PRRX{3.1.3.21} | 97712 | 18933 | PRRX1_MOUSE | P63013 | ||||
| PRRX2_MOUSE.H11MO.0.C |  |
MOUSE:Prrx2 | 7 | C | 0 | hTAATTR | HOCOMOCOv9 | Integrative | 65 | Paired-related HD factors{3.1.3} | PRRX{3.1.3.21} | 98218 | 20204 | PRRX2_MOUSE | Q06348 | ||||
| PTF1A_MOUSE.H11MO.0.A |  |
MOUSE:Ptf1a | 18 | A | 0 | vCAGCTGbbnbbbbYYhv | HOCOMOCOv11 | ChIP-Seq | 0.963 | 20 | 500 | Tal-related factors{1.2.3} | Twist-like factors{1.2.3.2} | 1328312 | 19213 | PTF1A_MOUSE | Q9QX98 | ||
| PTF1A_MOUSE.H11MO.1.A |  |
MOUSE:Ptf1a | 9 | A | 1 | RRCAGMTGK | HOCOMOCOv11 | ChIP-Seq | 0.93 | 20 | 503 | Tal-related factors{1.2.3} | Twist-like factors{1.2.3.2} | 1328312 | 19213 | PTF1A_MOUSE | Q9QX98 | ||
| PURA_MOUSE.H11MO.0.D |  |
MOUSE:Pura | 17 | D | 0 | RGWvKbKGnKGSMRKGK | HOCOMOCOv9 | Integrative | 13 | PUR{0.0.5} | Pur-alpha (PURA, PUR1){0.0.5.0.1} | 103079 | 19290 | PURA_MOUSE | P42669 | ||||
| RARA_MOUSE.H11MO.0.A |  |
MOUSE:Rara | 20 | A | 0 | RRdYYRdRAGKTCAAGGWSR | HOCOMOCOv11 | ChIP-Seq | 0.753 | 0.906 | 29 | 32 | 447 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Retinoic acid receptors (NR1B){2.1.2.1} | 97856 | 19401 | RARA_MOUSE | P11416 |
| RARA_MOUSE.H11MO.1.A |  |
MOUSE:Rara | 20 | A | 1 | vdddvdbbvdRAGKTCARRR | HOCOMOCOv11 | ChIP-Seq | 0.951 | 0.927 | 29 | 32 | 499 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Retinoic acid receptors (NR1B){2.1.2.1} | 97856 | 19401 | RARA_MOUSE | P11416 |
| RARA_MOUSE.H11MO.2.A |  |
MOUSE:Rara | 17 | A | 2 | RGKTSAnvvdvRSKKSR | HOCOMOCOv9 | Integrative | 0.806 | 0.844 | 29 | 32 | 122 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Retinoic acid receptors (NR1B){2.1.2.1} | 97856 | 19401 | RARA_MOUSE | P11416 |
| RARA_MOUSE.H11MO.3.A |  |
MOUSE:Rara | 12 | A | 3 | bhvAGGTCASvv | HOCOMOCOv11 | ChIP-Seq | 0.912 | 0.888 | 29 | 32 | 499 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Retinoic acid receptors (NR1B){2.1.2.1} | 97856 | 19401 | RARA_MOUSE | P11416 |
| RARB_MOUSE.H11MO.0.D |  |
MOUSE:Rarb | 11 | D | 0 | vAGGTCAnvRv | HOCOMOCOv9 | Integrative | 25 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Retinoic acid receptors (NR1B){2.1.2.1} | 97857 | 218772 | RARB_MOUSE | P22605 | ||||
| RARG_MOUSE.H11MO.0.C |  |
MOUSE:Rarg | 18 | C | 0 | RRGTCAvvvhSvMMbddv | HOCOMOCOv11 | ChIP-Seq | 0.827 | 12 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Retinoic acid receptors (NR1B){2.1.2.1} | 97858 | 19411 | RARG_MOUSE | P18911 | ||
| RARG_MOUSE.H11MO.1.C |  |
MOUSE:Rarg | 8 | C | 1 | vAGGKCAn | HOCOMOCOv11 | ChIP-Seq | 0.815 | 12 | 394 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Retinoic acid receptors (NR1B){2.1.2.1} | 97858 | 19411 | RARG_MOUSE | P18911 | ||
| RELB_MOUSE.H11MO.0.C |  |
MOUSE:Relb | 12 | C | 0 | vRGGGMTTTSMh | HOCOMOCOv9 | Integrative | 28 | NF-kappaB-related factors{6.1.1} | NF-kappaB p65 subunit-like factors{6.1.1.2} | 103289 | 19698 | RELB_MOUSE | Q04863 | ||||
| REL_MOUSE.H11MO.0.A |  |
MOUSE:Rel | 15 | A | 0 | dvhdGGGRAWKbMSW | HOCOMOCOv11 | ChIP-Seq | 0.921 | 11 | 178 | NF-kappaB-related factors{6.1.1} | NF-kappaB p65 subunit-like factors{6.1.1.2} | 97897 | REL_MOUSE | P15307 | |||
| REST_MOUSE.H11MO.0.A |  |
MOUSE:Rest | 22 | A | 0 | dYCAGSACCdhGGACAGMdSMh | HOCOMOCOv11 | ChIP-Seq | 0.99 | 0.982 | 147 | 33 | 378 | Factors with multiple dispersed zinc fingers{2.3.4} | unclassified{2.3.4.0} | 104897 | 19712 | REST_MOUSE | Q8VIG1 |
| RFX1_MOUSE.H11MO.0.A |  |
MOUSE:Rfx1 | 22 | A | 0 | bbbbSbGTTGCCWdGGvvMSvv | HOCOMOCOv10 | ChIP-Seq | 0.712 | 0.995 | 3 | 7 | 457 | RFX-related factors{3.3.3} | RFX1 (EF-C){3.3.3.0.1} | 105982 | 19724 | RFX1_MOUSE | P48377 |
| RFX1_MOUSE.H11MO.1.A |  |
MOUSE:Rfx1 | 11 | A | 1 | bGTTGCCWdGR | HOCOMOCOv11 | ChIP-Seq | 0.574 | 0.994 | 3 | 7 | 500 | RFX-related factors{3.3.3} | RFX1 (EF-C){3.3.3.0.1} | 105982 | 19724 | RFX1_MOUSE | P48377 |
| RFX2_MOUSE.H11MO.0.A |  |
MOUSE:Rfx2 | 22 | A | 0 | vbSnGTYRYYdKGGvRMYvdvn | HOCOMOCOv11 | ChIP-Seq | 0.937 | 0.997 | 7 | 7 | 132 | RFX-related factors{3.3.3} | RFX2{3.3.3.0.2} | 106583 | 19725 | RFX2_MOUSE | P48379 |
| RFX2_MOUSE.H11MO.1.A |  |
MOUSE:Rfx2 | 11 | A | 1 | bGTTGCCWdGG | HOCOMOCOv11 | ChIP-Seq | 0.96 | 0.997 | 7 | 7 | 500 | RFX-related factors{3.3.3} | RFX2{3.3.3.0.2} | 106583 | 19725 | RFX2_MOUSE | P48379 |
| RFX3_MOUSE.H11MO.0.C |  |
MOUSE:Rfx3 | 16 | C | 0 | SbGTTGCCWKGGvRAC | HOCOMOCOv11 | ChIP-Seq | 0.989 | 2 | 351 | RFX-related factors{3.3.3} | RFX3{3.3.3.0.3} | 106582 | 19726 | RFX3_MOUSE | P48381 | ||
| RFX3_MOUSE.H11MO.1.C |  |
MOUSE:Rfx3 | 10 | C | 1 | bGTTGCCWKG | HOCOMOCOv11 | ChIP-Seq | 0.997 | 2 | 500 | RFX-related factors{3.3.3} | RFX3{3.3.3.0.3} | 106582 | 19726 | RFX3_MOUSE | P48381 | ||
| RFX6_MOUSE.H11MO.0.C |  |
MOUSE:Rfx6 | 11 | C | 0 | bdGYWKCYnKG | HOCOMOCOv11 | ChIP-Seq | 0.998 | 3 | 501 | RFX-related factors{3.3.3} | RFX6 (RFXDC-1){3.3.3.0.6} | 2445208 | 320995 | RFX6_MOUSE | Q8C7R7 | ||
| RFX6_MOUSE.H11MO.1.C |  |
MOUSE:Rfx6 | 14 | C | 1 | dGYTGCYdKGvdvv | HOCOMOCOv11 | ChIP-Seq | 0.979 | 3 | 502 | RFX-related factors{3.3.3} | RFX6 (RFXDC-1){3.3.3.0.6} | 2445208 | 320995 | RFX6_MOUSE | Q8C7R7 | ||
| RORA_MOUSE.H11MO.0.C |  |
MOUSE:Rora | 13 | C | 0 | vhbdRGGbCAvdR | HOCOMOCOv11 | ChIP-Seq | 0.682 | 0.788 | 2 | 5 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | ROR (NR1F){2.1.2.6} | 104661 | 19883 | RORA_MOUSE | P51448 |
| RORG_MOUSE.H11MO.0.B |  |
MOUSE:Rorc | 12 | B | 0 | vdRWSTRRGKCA | HOCOMOCOv11 | ChIP-Seq | 0.91 | 19 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | ROR (NR1F){2.1.2.6} | 104856 | 19885 | RORG_MOUSE | P51450 | ||
| RREB1_MOUSE.H11MO.0.D |  |
MOUSE:Rreb1 | 22 | D | 0 | dbSKbdGKKGKTGbTKKGKGGd | HOCOMOCOv9 | Integrative | 26 | Factors with multiple dispersed zinc fingers{2.3.4} | unclassified{2.3.4.0} | 2443664 | 68750 | RREB1_MOUSE | Q3UH06 | ||||
| RUNX1_MOUSE.H11MO.0.A |  |
MOUSE:Runx1 | 12 | A | 0 | bbYTGTGGTYWb | HOCOMOCOv11 | ChIP-Seq | 0.963 | 0.984 | 115 | 63 | 500 | Runt-related factors{6.4.1} | Runx1 (PEBP2alphaB, CBF-alpha2, AML-1){6.4.1.0.2} | 99852 | RUNX1_MOUSE | Q03347 | |
| RUNX2_MOUSE.H11MO.0.A |  |
MOUSE:Runx2 | 12 | A | 0 | bbYTGTGGYYdb | HOCOMOCOv11 | ChIP-Seq | 0.837 | 0.927 | 3 | 25 | 501 | Runt-related factors{6.4.1} | Runx2 (PEBP2alphaA, CBF-alpha1){6.4.1.0.1} | 99829 | 12393 | RUNX2_MOUSE | Q08775 |
| RUNX3_MOUSE.H11MO.0.A |  |
MOUSE:Runx3 | 12 | A | 0 | bbhTGTGGTYdb | HOCOMOCOv11 | ChIP-Seq | 0.996 | 0.975 | 2 | 27 | 500 | Runt-related factors{6.4.1} | Runx3 (PEBP2alphaC, CBF-alpha3, AML-2){6.4.1.0.3} | 102672 | RUNX3_MOUSE | Q64131 | |
| RXRA_MOUSE.H11MO.0.A |  |
MOUSE:Rxra | 21 | A | 0 | ddvdbvvdRAGKTCARGGbYR | HOCOMOCOv11 | ChIP-Seq | 0.782 | 0.943 | 52 | 158 | 499 | RXR-related receptors (NR2){2.1.3} | Retinoid X receptors (NR2B){2.1.3.1} | 98214 | 20181 | RXRA_MOUSE | P28700 |
| RXRA_MOUSE.H11MO.1.A |  |
MOUSE:Rxra | 22 | A | 1 | vdddvdbvvdRAGKTCAvRRhb | HOCOMOCOv11 | ChIP-Seq | 0.734 | 0.97 | 52 | 158 | 500 | RXR-related receptors (NR2){2.1.3} | Retinoid X receptors (NR2B){2.1.3.1} | 98214 | 20181 | RXRA_MOUSE | P28700 |
| RXRA_MOUSE.H11MO.2.A |  |
MOUSE:Rxra | 10 | A | 2 | RRGGTCAdRR | HOCOMOCOv11 | ChIP-Seq | 0.784 | 0.963 | 52 | 158 | 365 | RXR-related receptors (NR2){2.1.3} | Retinoid X receptors (NR2B){2.1.3.1} | 98214 | 20181 | RXRA_MOUSE | P28700 |
| RXRB_MOUSE.H11MO.0.C |  |
MOUSE:Rxrb | 10 | C | 0 | WSAGGTCAbd | HOCOMOCOv9 | Integrative | 30 | RXR-related receptors (NR2){2.1.3} | Retinoid X receptors (NR2B){2.1.3.1} | 98215 | 20182 | RXRB_MOUSE | P28704 | ||||
| RXRG_MOUSE.H11MO.0.B |  |
MOUSE:Rxrg | 22 | B | 0 | vddvdbbvdRAGKTCAAGGYYW | HOCOMOCOv11 | ChIP-Seq | 0.952 | 7 | 281 | RXR-related receptors (NR2){2.1.3} | Retinoid X receptors (NR2B){2.1.3.1} | 98216 | 20183 | RXRG_MOUSE | P28705 | ||
| RXRG_MOUSE.H11MO.1.B |  |
MOUSE:Rxrg | 10 | B | 1 | RAGKTCARRS | HOCOMOCOv11 | ChIP-Seq | 0.946 | 7 | 501 | RXR-related receptors (NR2){2.1.3} | Retinoid X receptors (NR2B){2.1.3.1} | 98216 | 20183 | RXRG_MOUSE | P28705 | ||
| SALL1_MOUSE.H11MO.0.D |  |
MOUSE:Sall1 | 17 | D | 0 | ddvRGGSWGKGRbvddv | HOCOMOCOv11 | ChIP-Seq | 0.799 | 3 | 500 | Factors with multiple dispersed zinc fingers{2.3.4} | Sal-like factors{2.3.4.3} | 1889585 | SALL1_MOUSE | Q9ER74 | |||
| SALL4_MOUSE.H11MO.0.A |  |
MOUSE:Sall4 | 10 | A | 0 | dvdGGGWGKG | HOCOMOCOv11 | ChIP-Seq | 0.724 | 0.924 | 1 | 21 | 500 | Factors with multiple dispersed zinc fingers{2.3.4} | Sal-like factors{2.3.4.3} | 2139360 | 99377 | SALL4_MOUSE | Q8BX22 |
| SIX2_MOUSE.H11MO.0.A |  |
MOUSE:Six2 | 14 | A | 0 | KGWAAYhYRAbMYb | HOCOMOCOv11 | ChIP-Seq | 0.963 | 0.946 | 18 | 9 | 500 | HD-SINE factors{3.1.6} | SIX1-like factors{3.1.6.1} | 102778 | 20472 | SIX2_MOUSE | Q62232 |
| SIX4_MOUSE.H11MO.0.C |  |
MOUSE:Six4 | 14 | C | 0 | dGdMAbhdSAvhbb | HOCOMOCOv11 | ChIP-Seq | 0.912 | 4 | 487 | HD-SINE factors{3.1.6} | SIX4-like factors{3.1.6.3} | 106034 | 20474 | SIX4_MOUSE | Q61321 | ||
| SMAD1_MOUSE.H11MO.0.D |  |
MOUSE:Smad1 | 12 | D | 0 | dGCCTSTCTGYM | HOCOMOCOv9 | Integrative | 12 | SMAD factors{7.1.1} | Regulatory Smads (R-Smad){7.1.1.1} | 109452 | 17125 | SMAD1_MOUSE | P70340 | ||||
| SMAD2_MOUSE.H11MO.0.A |  |
MOUSE:Smad2 | 9 | A | 0 | dvTGYCTGb | HOCOMOCOv11 | ChIP-Seq | 0.803 | 0.885 | 17 | 26 | 500 | SMAD factors{7.1.1} | Regulatory Smads (R-Smad){7.1.1.1} | 108051 | 17126 | SMAD2_MOUSE | Q62432 |
| SMAD3_MOUSE.H11MO.0.B |  |
MOUSE:Smad3 | 12 | B | 0 | vbYTShCWSCWb | HOCOMOCOv11 | ChIP-Seq | 0.806 | 0.707 | 34 | 30 | 500 | SMAD factors{7.1.1} | Regulatory Smads (R-Smad){7.1.1.1} | 1201674 | 17127 | SMAD3_MOUSE | Q8BUN5 |
| SMAD4_MOUSE.H11MO.0.A |  |
MOUSE:Smad4 | 12 | A | 0 | vWGbCTvnChSh | HOCOMOCOv11 | ChIP-Seq | 0.835 | 0.873 | 24 | 3 | 489 | SMAD factors{7.1.1} | Co-activating Smads (Co-Smad){7.1.1.2} | 894293 | 17128 | SMAD4_MOUSE | P97471 |
| SMCA5_MOUSE.H11MO.0.C |  |
MOUSE:Smarca5 | 15 | C | 0 | WGGMvKnSRWTGGAR | HOCOMOCOv11 | ChIP-Seq | 0.754 | 0.741 | 3 | 2 | 419 | Myb/SANT domain factors{3.5.1} | SMARCA-like factors{3.5.1.5} | 1935129 | 93762 | SMCA5_MOUSE | Q91ZW3 |
| SNAI1_MOUSE.H11MO.0.C |  |
MOUSE:Snai1 | 8 | C | 0 | vCAGGTGb | HOCOMOCOv9 | Integrative | 5 | More than 3 adjacent zinc finger factors{2.3.3} | Snail-like factors{2.3.3.2} | 98330 | 20613 | SNAI1_MOUSE | Q02085 | ||||
| SNAI2_MOUSE.H11MO.0.A |  |
MOUSE:Snai2 | 11 | A | 0 | dRCAGGTGYnb | HOCOMOCOv11 | ChIP-Seq | 0.974 | 0.875 | 11 | 3 | 494 | More than 3 adjacent zinc finger factors{2.3.3} | Snail-like factors{2.3.3.2} | 1096393 | 20583 | SNAI2_MOUSE | P97469 |
| SOX10_MOUSE.H11MO.0.B |  |
MOUSE:Sox10 | 16 | B | 0 | vARdRvvnbYWTTGTb | HOCOMOCOv11 | ChIP-Seq | 0.91 | 0.763 | 3 | 3 | 500 | 98358 | 20665 | SOX10_MOUSE | Q04888 | ||
| SOX10_MOUSE.H11MO.1.A |  |
MOUSE:Sox10 | 11 | A | 1 | bbCWTTGTbbn | HOCOMOCOv11 | ChIP-Seq | 0.82 | 0.831 | 3 | 3 | 505 | 98358 | 20665 | SOX10_MOUSE | Q04888 | ||
| SOX13_MOUSE.H11MO.0.D |  |
MOUSE:Sox13 | 8 | D | 0 | nATTGTKn | HOCOMOCOv9 | Integrative | 5 | SOX-related factors{4.1.1} | Group D{4.1.1.4} | 98361 | 20668 | SOX13_MOUSE | Q04891 | ||||
| SOX15_MOUSE.H11MO.0.D |  |
MOUSE:Sox15 | 7 | D | 0 | CWTTGTT | HOCOMOCOv9 | Integrative | 8 | SOX-related factors{4.1.1} | Group G{4.1.1.7} | 98363 | 20670 | SOX15_MOUSE | P43267 | ||||
| SOX17_MOUSE.H11MO.0.D |  |
MOUSE:Sox17 | 11 | D | 0 | bCCATYKTbbb | HOCOMOCOv11 | ChIP-Seq | 0.835 | 4 | 500 | SOX-related factors{4.1.1} | Group F{4.1.1.6} | 107543 | 20671 | SOX17_MOUSE | Q61473 | ||
| SOX18_MOUSE.H11MO.0.D |  |
MOUSE:Sox18 | 19 | D | 0 | SMWhnMATTGTYbWnnYYC | HOCOMOCOv9 | Integrative | 8 | SOX-related factors{4.1.1} | Group F{4.1.1.6} | 103559 | 20672 | SOX18_MOUSE | P43680 | ||||
| SOX2_MOUSE.H11MO.0.A |  |
MOUSE:Sox2 | 16 | A | 0 | bbYWTTSTnWTKYWdW | HOCOMOCOv11 | ChIP-Seq | 0.955 | 0.958 | 67 | 109 | 499 | SOX-related factors{4.1.1} | Group B{4.1.1.2} | 98364 | 20674 | SOX2_MOUSE | P48432 |
| SOX2_MOUSE.H11MO.1.A |  |
MOUSE:Sox2 | 13 | A | 1 | hbYYWTTGTbhbb | HOCOMOCOv11 | ChIP-Seq | 0.944 | 0.926 | 67 | 109 | 500 | SOX-related factors{4.1.1} | Group B{4.1.1.2} | 98364 | 20674 | SOX2_MOUSE | P48432 |
| SOX3_MOUSE.H11MO.0.C |  |
MOUSE:Sox3 | 11 | C | 0 | bYCTTTGTbnb | HOCOMOCOv11 | ChIP-Seq | 0.559 | 0.914 | 2 | 4 | 344 | SOX-related factors{4.1.1} | Group B{4.1.1.2} | 98365 | SOX3_MOUSE | P53784 | |
| SOX4_MOUSE.H11MO.0.A |  |
MOUSE:Sox4 | 12 | A | 0 | bYCTTTGTYYbb | HOCOMOCOv11 | ChIP-Seq | 0.64 | 0.978 | 2 | 7 | 500 | SOX-related factors{4.1.1} | Group C{4.1.1.3} | 98366 | 20677 | SOX4_MOUSE | Q06831 |
| SOX5_MOUSE.H11MO.0.C |  |
MOUSE:Sox5 | 8 | C | 0 | bATTGTTh | HOCOMOCOv9 | Integrative | 69 | SOX-related factors{4.1.1} | Group D{4.1.1.4} | 98367 | 20678 | SOX5_MOUSE | P35710 | ||||
| SOX9_MOUSE.H11MO.0.A |  |
MOUSE:Sox9 | 16 | A | 0 | AAdRvnnbYWTTGTKY | HOCOMOCOv11 | ChIP-Seq | 0.718 | 0.981 | 6 | 25 | 500 | SOX-related factors{4.1.1} | Group E{4.1.1.5} | 98371 | 20682 | SOX9_MOUSE | Q04887 |
| SOX9_MOUSE.H11MO.1.A |  |
MOUSE:Sox9 | 11 | A | 1 | bCWYTGTKYhb | HOCOMOCOv11 | ChIP-Seq | 0.679 | 0.93 | 6 | 25 | 499 | SOX-related factors{4.1.1} | Group E{4.1.1.5} | 98371 | 20682 | SOX9_MOUSE | Q04887 |
| SP1_MOUSE.H11MO.0.A |  |
MOUSE:Sp1 | 24 | A | 0 | SSvSRvRSvRRRGGMGGRGMvKSR | HOCOMOCOv11 | ChIP-Seq | 0.999 | 0.988 | 53 | 11 | 489 | Three-zinc finger Krüppel-related factors{2.3.1} | Sp1-like factors{2.3.1.1} | 98372 | 20683 | SP1_MOUSE | O89090 |
| SP1_MOUSE.H11MO.1.A |  |
MOUSE:Sp1 | 11 | A | 1 | RKGGGMGGRRv | HOCOMOCOv9 | Integrative | 0.996 | 0.986 | 53 | 11 | 3614 | Three-zinc finger Krüppel-related factors{2.3.1} | Sp1-like factors{2.3.1.1} | 98372 | 20683 | SP1_MOUSE | O89090 |
| SP2_MOUSE.H11MO.0.B |  |
MOUSE:Sp2 | 22 | B | 0 | SvvvvRRRGGCGGRRSbnvvSv | HOCOMOCOv10 | ChIP-Seq | 0.972 | 16 | 499 | Three-zinc finger Krüppel-related factors{2.3.1} | Sp1-like factors{2.3.1.1} | 1926162 | 78912 | SP2_MOUSE | Q9D2H6 | ||
| SP2_MOUSE.H11MO.1.C |  |
MOUSE:Sp2 | 12 | C | 1 | dRRGGGCGGGRM | HOCOMOCOv11 | ChIP-Seq | 0.893 | 16 | 498 | Three-zinc finger Krüppel-related factors{2.3.1} | Sp1-like factors{2.3.1.1} | 1926162 | 78912 | SP2_MOUSE | Q9D2H6 | ||
| SP3_MOUSE.H11MO.0.B |  |
MOUSE:Sp3 | 20 | B | 0 | vvvvvKGGGMGGRSvbdvvv | HOCOMOCOv9 | Integrative | 415 | Three-zinc finger Krüppel-related factors{2.3.1} | Sp1-like factors{2.3.1.1} | 1277166 | 20687 | SP3_MOUSE | O70494 | ||||
| SP4_MOUSE.H11MO.0.B |  |
MOUSE:Sp4 | 20 | B | 0 | vvSvdRRRGGMGGRGMhdvv | HOCOMOCOv11 | ChIP-Seq | 0.983 | 5 | 499 | Three-zinc finger Krüppel-related factors{2.3.1} | Sp1-like factors{2.3.1.1} | 107595 | SP4_MOUSE | Q62445 | |||
| SP4_MOUSE.H11MO.1.B |  |
MOUSE:Sp4 | 16 | B | 1 | RvRRRGGGMGGRGvnd | HOCOMOCOv11 | ChIP-Seq | 0.983 | 5 | 496 | Three-zinc finger Krüppel-related factors{2.3.1} | Sp1-like factors{2.3.1.1} | 107595 | SP4_MOUSE | Q62445 | |||
| SP5_MOUSE.H11MO.0.C |  |
MOUSE:Sp5 | 24 | C | 0 | vdddRRRvRRvRvdRRGGGMGGRG | HOCOMOCOv11 | ChIP-Seq | 0.956 | 3 | 499 | Three-zinc finger Krüppel-related factors{2.3.1} | Sp1-like factors{2.3.1.1} | 1927715 | 64406 | SP5_MOUSE | Q9JHX2 | ||
| SP5_MOUSE.H11MO.1.C |  |
MOUSE:Sp5 | 13 | C | 1 | dvdvdGGGMGGRG | HOCOMOCOv11 | ChIP-Seq | 0.94 | 3 | 458 | Three-zinc finger Krüppel-related factors{2.3.1} | Sp1-like factors{2.3.1.1} | 1927715 | 64406 | SP5_MOUSE | Q9JHX2 | ||
| SP7_MOUSE.H11MO.0.A |  |
MOUSE:Sp7 | 14 | A | 0 | vhAvSMYRYMATTW | HOCOMOCOv11 | ChIP-Seq | 0.976 | 19 | 500 | Three-zinc finger Krüppel-related factors{2.3.1} | Sp1-like factors{2.3.1.1} | 2153568 | 170574 | SP7_MOUSE | Q8VI67 | ||
| SPI1_MOUSE.H11MO.0.A |  |
MOUSE:Spi1 | 15 | A | 0 | vvRRWGRGGAAGTGn | HOCOMOCOv11 | ChIP-Seq | 0.999 | 0.997 | 80 | 204 | 482 | Ets-related factors{3.5.2} | Spi-like factors{3.5.2.5} | 98282 | 20375 | SPI1_MOUSE | P17433 |
| SPIB_MOUSE.H11MO.0.A |  |
MOUSE:Spib | 17 | A | 0 | RAAWGRGGAAGTGRvWn | HOCOMOCOv11 | ChIP-Seq | 0.996 | 0.977 | 12 | 3 | 458 | Ets-related factors{3.5.2} | Spi-like factors{3.5.2.5} | 892986 | 272382 | SPIB_MOUSE | O35906 |
| SPZ1_MOUSE.H11MO.0.D |  |
MOUSE:Spz1 | 16 | D | 0 | nSRSKKTTMCMbhbGn | HOCOMOCOv9 | Integrative | 11 | 1930801 | 79401 | SPZ1_MOUSE | Q99MY0 | ||||||
| SRBP1_MOUSE.H11MO.0.B |  |
MOUSE:Srebf1 | 11 | B | 0 | vRTSRSSWGWb | HOCOMOCOv9 | Integrative | 0.893 | 0.735 | 5 | 23 | 2191 | bHLH-ZIP factors{1.2.6} | SREBP factors{1.2.6.3} | 107606 | 20787 | SRBP1_MOUSE | Q9WTN3 |
| SRBP2_MOUSE.H11MO.0.B |  |
MOUSE:Srebf2 | 13 | B | 0 | vvvWGGvSWGRnb | HOCOMOCOv9 | Integrative | 2363 | bHLH-ZIP factors{1.2.6} | SREBP factors{1.2.6.3} | 107585 | 20788 | SRBP2_MOUSE | Q3U1N2 | ||||
| SRF_MOUSE.H11MO.0.A |  |
MOUSE:Srf | 17 | A | 0 | hKbCCWTWTWWGGbhdb | HOCOMOCOv11 | ChIP-Seq | 0.95 | 0.986 | 46 | 37 | 374 | Responders to external signals (SRF/RLM1){5.1.2} | SRF{5.1.2.0.1} | 106658 | 20807 | SRF_MOUSE | Q9JM73 |
| SRY_MOUSE.H11MO.0.B |  |
MOUSE:Sry | 9 | B | 0 | bWTTGTWWh | HOCOMOCOv9 | Integrative | 108 | SOX-related factors{4.1.1} | Group A{4.1.1.1} | 98660 | 21674 | SRY_MOUSE | Q05738 | ||||
| STA5A_MOUSE.H11MO.0.A |  |
MOUSE:Stat5a | 10 | A | 0 | TTCYARGAAv | HOCOMOCOv11 | ChIP-Seq | 0.928 | 0.992 | 24 | 127 | 500 | STAT factors{6.2.1} | STAT5A{6.2.1.0.5} | 103036 | 20850 | STA5A_MOUSE | P42230 |
| STA5B_MOUSE.H11MO.0.A |  |
MOUSE:Stat5b | 12 | A | 0 | nTTCYTRGAAdh | HOCOMOCOv11 | ChIP-Seq | 0.965 | 0.957 | 16 | 31 | 501 | STAT factors{6.2.1} | STAT5B{6.2.1.0.6} | 103035 | 20851 | STA5B_MOUSE | P42232 |
| STAT1_MOUSE.H11MO.0.A |  |
MOUSE:Stat1 | 22 | A | 0 | dRRRAAhdGAAAvhvMvnvhdd | HOCOMOCOv11 | ChIP-Seq | 0.989 | 0.958 | 37 | 128 | 500 | STAT factors{6.2.1} | STAT1{6.2.1.0.1} | 103063 | STAT1_MOUSE | P42225 | |
| STAT1_MOUSE.H11MO.1.A |  |
MOUSE:Stat1 | 21 | A | 1 | bvdhTKMhRGGAARhvRRvhv | HOCOMOCOv11 | ChIP-Seq | 0.976 | 0.951 | 37 | 128 | 500 | STAT factors{6.2.1} | STAT1{6.2.1.0.1} | 103063 | STAT1_MOUSE | P42225 | |
| STAT2_MOUSE.H11MO.0.A |  |
MOUSE:Stat2 | 19 | A | 0 | dvRRAAhhGAAASYdRRRv | HOCOMOCOv11 | ChIP-Seq | 0.98 | 0.994 | 11 | 8 | 500 | STAT factors{6.2.1} | STAT2{6.2.1.0.2} | 103039 | STAT2_MOUSE | Q9WVL2 | |
| STAT3_MOUSE.H11MO.0.A |  |
MOUSE:Stat3 | 12 | A | 0 | bdYTTCCMRKMA | HOCOMOCOv11 | ChIP-Seq | 0.94 | 0.941 | 36 | 82 | 500 | STAT factors{6.2.1} | STAT3{6.2.1.0.3} | 103038 | 20848 | STAT3_MOUSE | P42227 |
| STAT4_MOUSE.H11MO.0.A |  |
MOUSE:Stat4 | 10 | A | 0 | YTTCYhdKMM | HOCOMOCOv11 | ChIP-Seq | 0.932 | 0.914 | 7 | 10 | 475 | STAT factors{6.2.1} | STAT4{6.2.1.0.4} | 103062 | 20849 | STAT4_MOUSE | P42228 |
| STAT6_MOUSE.H11MO.0.A |  |
MOUSE:Stat6 | 14 | A | 0 | bTTCYhvRGAAvhv | HOCOMOCOv11 | ChIP-Seq | 0.751 | 0.932 | 19 | 24 | 407 | STAT factors{6.2.1} | STAT6{6.2.1.0.7} | 103034 | 20852 | STAT6_MOUSE | P52633 |
| STF1_MOUSE.H11MO.0.B |  |
MOUSE:Nr5a1 | 11 | B | 0 | bbYCAAGGYCW | HOCOMOCOv11 | ChIP-Seq | 0.759 | 0.898 | 1 | 2 | 500 | FTZ-F1-related receptors (NR5){2.1.5} | FTZ-F1 (SF-1) (NR5A1){2.1.5.0.1} | 1346833 | 26423 | STF1_MOUSE | P33242 |
| SUH_MOUSE.H11MO.0.A |  |
MOUSE:Rbpj | 11 | A | 0 | bbbTGRGAAvv | HOCOMOCOv11 | ChIP-Seq | 0.939 | 0.973 | 25 | 6 | 500 | CSL-related factors{6.1.4} | M{6.1.4.1} | 96522 | 19664 | SUH_MOUSE | P31266 |
| TAF1_MOUSE.H11MO.0.A |  |
MOUSE:Taf1 | 14 | A | 0 | vRRRAWGGMRRhbv | HOCOMOCOv11 | ChIP-Seq | 0.813 | 0.956 | 61 | 2 | 500 | TCF-7-related factors{4.1.3} | TAF-1 (TAF-2A, TAF(II)250) [1]{4.1.3.0.5} | 1336878 | 270627 | TAF1_MOUSE | Q80UV9 |
| TAL1_MOUSE.H11MO.0.A |  |
MOUSE:Tal1 | 20 | A | 0 | bYKbnnnnndvAGATAASvv | HOCOMOCOv11 | ChIP-Seq | 0.965 | 0.978 | 68 | 76 | 500 | Tal-related factors{1.2.3} | Tal / HEN-like factors{1.2.3.1} | 98480 | 21349 | TAL1_MOUSE | P22091 |
| TBP_MOUSE.H11MO.0.A |  |
MOUSE:Tbp | 10 | A | 0 | vWATAAAWRv | HOCOMOCOv11 | ChIP-Seq | 0.788 | 0.851 | 21 | 45 | 500 | TBP-related factors{8.1.1} | TBP{8.1.1.0.1} | 101838 | 21374 | TBP_MOUSE | P29037 |
| TBX20_MOUSE.H11MO.0.C |  |
MOUSE:Tbx20 | 14 | C | 0 | ddSTGnTGAMARvn | HOCOMOCOv11 | ChIP-Seq | 0.883 | 4 | 500 | TBX1-related factors{6.5.3} | TBX20{6.5.3.0.5} | 1888496 | 57246 | TBX20_MOUSE | Q9ES03 | ||
| TBX21_MOUSE.H11MO.0.A |  |
MOUSE:Tbx21 | 12 | A | 0 | dddKGTGdSWdd | HOCOMOCOv11 | ChIP-Seq | 0.804 | 0.867 | 7 | 17 | 507 | TBrain-related factors{6.5.2} | TBX21 (T-bet){6.5.2.0.3} | 1888984 | 57765 | TBX21_MOUSE | Q9JKD8 |
| TBX2_MOUSE.H11MO.0.D |  |
MOUSE:Tbx2 | 25 | D | 0 | hbMhWhChbAGGTGTGAnRWKnSMn | HOCOMOCOv9 | Integrative | 9 | TBX2-related factors{6.5.4} | TBX2{6.5.4.0.1} | 98494 | 21385 | TBX2_MOUSE | Q60707 | ||||
| TBX3_MOUSE.H11MO.0.B |  |
MOUSE:Tbx3 | 9 | B | 0 | RRGGTGbbR | HOCOMOCOv11 | ChIP-Seq | 0.756 | 0.955 | 2 | 3 | 114 | TBX2-related factors{6.5.4} | TBX3{6.5.4.0.2} | 98495 | 21386 | TBX3_MOUSE | P70324 |
| TBX5_MOUSE.H11MO.0.D |  |
MOUSE:Tbx5 | 8 | D | 0 | ARGTGKSW | HOCOMOCOv9 | Integrative | 38 | TBX2-related factors{6.5.4} | TBX5{6.5.4.0.4} | 102541 | 21388 | TBX5_MOUSE | P70326 | ||||
| TCF7_MOUSE.H11MO.0.A |  |
MOUSE:Tcf7 | 16 | A | 0 | vvRvWdSAAARvvvvv | HOCOMOCOv11 | ChIP-Seq | 0.78 | 0.947 | 7 | 19 | 500 | TCF-7-related factors{4.1.3} | TCF-7 (TCF-1) [1]{4.1.3.0.1} | 98507 | 21414 | TCF7_MOUSE | Q00417 |
| TEAD1_MOUSE.H11MO.0.A |  |
MOUSE:Tead1 | 11 | A | 0 | RMATTCCWRSv | HOCOMOCOv11 | ChIP-Seq | 0.949 | 0.95 | 14 | 5 | 500 | TEF-1-related factors{3.6.1} | TEF-1 (TEAD-1, TCF-13){3.6.1.0.1} | 101876 | 21676 | TEAD1_MOUSE | P30051 |
| TEAD2_MOUSE.H11MO.0.C |  |
MOUSE:Tead2 | 12 | C | 0 | hbRMATTCCWRv | HOCOMOCOv11 | ChIP-Seq | 0.519 | 0.936 | 2 | 2 | 308 | TEF-1-related factors{3.6.1} | TEF-4 (TEAD-2){3.6.1.0.3} | 104904 | 21677 | TEAD2_MOUSE | P48301 |
| TEAD3_MOUSE.H11MO.0.D |  |
MOUSE:Tead3 | 15 | D | 0 | nAWATTYCTGhhbYh | HOCOMOCOv9 | Integrative | 12 | TEF-1-related factors{3.6.1} | TEF-5 (TEAD-3, TEAD5){3.6.1.0.4} | 109241 | 21678 | TEAD3_MOUSE | P70210 | ||||
| TEAD4_MOUSE.H11MO.0.A |  |
MOUSE:Tead4 | 15 | A | 0 | RMATTCCWvdvdYdb | HOCOMOCOv11 | ChIP-Seq | 0.963 | 0.946 | 57 | 10 | 500 | TEF-1-related factors{3.6.1} | TEF-3 (TEAD-4, TCF-13L1, Tefr1){3.6.1.0.2} | 106907 | 21679 | TEAD4_MOUSE | Q62296 |
| TEF_MOUSE.H11MO.0.D |  |
MOUSE:Tef | 14 | D | 0 | TGdTKAYRTMdvYd | HOCOMOCOv9 | Integrative | 14 | C/EBP-related{1.1.8} | PAR factors{1.1.8.2} | 98663 | 21685 | TEF_MOUSE | Q9JLC6 | ||||
| TF2L1_MOUSE.H11MO.0.C |  |
MOUSE:Tfcp2l1 | 19 | C | 0 | ddCYRGYYYhddCYRGhYb | HOCOMOCOv11 | ChIP-Seq | 0.993 | 3 | 491 | CP2-related factors{6.7.2} | CP2-L1 (LBP-9, CRTR-1){6.7.2.0.2} | 2444691 | 81879 | TF2L1_MOUSE | Q3UNW5 | ||
| TF2L1_MOUSE.H11MO.1.C |  |
MOUSE:Tfcp2l1 | 10 | C | 1 | dRRCYGGYYb | HOCOMOCOv11 | ChIP-Seq | 0.962 | 3 | 502 | CP2-related factors{6.7.2} | CP2-L1 (LBP-9, CRTR-1){6.7.2.0.2} | 2444691 | 81879 | TF2L1_MOUSE | Q3UNW5 | ||
| TF65_MOUSE.H11MO.0.A |  |
MOUSE:Rela | 11 | A | 0 | KGGRvTTTCCM | HOCOMOCOv11 | ChIP-Seq | 0.974 | 0.931 | 188 | 99 | 500 | NF-kappaB-related factors{6.1.1} | NF-kappaB p65 subunit-like factors{6.1.1.2} | 103290 | 19697 | TF65_MOUSE | Q04207 |
| TF7L1_MOUSE.H11MO.0.A |  |
MOUSE:Tcf7l1 | 11 | A | 0 | bbCTTTSAWSW | HOCOMOCOv11 | ChIP-Seq | 0.867 | 0.865 | 10 | 10 | 500 | TCF-7-related factors{4.1.3} | TCF-7L1 (TCF-3) [1]{4.1.3.0.2} | 1202876 | 21415 | TF7L1_MOUSE | Q9Z1J1 |
| TF7L2_MOUSE.H11MO.0.A |  |
MOUSE:Tcf7l2 | 13 | A | 0 | bbCTTTGATSYbb | HOCOMOCOv11 | ChIP-Seq | 0.962 | 0.888 | 46 | 10 | 500 | TCF-7-related factors{4.1.3} | TCF-7L2 (TCF-4) [1]{4.1.3.0.3} | 1202879 | 21416 | TF7L2_MOUSE | Q924A0 |
| TFCP2_MOUSE.H11MO.0.D |  |
MOUSE:Tfcp2 | 16 | D | 0 | SSCWGvnMdvRCMRGM | HOCOMOCOv9 | Integrative | 13 | CP2-related factors{6.7.2} | CP2 (LSF, SEF){6.7.2.0.1} | 98509 | 21422 | TFCP2_MOUSE | Q9ERA0 | ||||
| TFDP1_MOUSE.H11MO.0.D |  |
MOUSE:Tfdp1 | 14 | D | 0 | vvddSGCGGGAMdn | HOCOMOCOv11 | ChIP-Seq | 0.871 | 3 | 500 | E2F-related factors{3.3.2} | Dp-1{3.3.2.2} | 101934 | 21781 | TFDP1_MOUSE | Q08639 | ||
| TFE2_MOUSE.H11MO.0.A |  |
MOUSE:Tcf3 | 11 | A | 0 | vMCASCTGShn | HOCOMOCOv11 | ChIP-Seq | 0.926 | 0.977 | 9 | 60 | 503 | E2A-related factors{1.2.1} | E2A (TCF-3, ITF-1){1.2.1.0.1} | 98510 | 21423 | TFE2_MOUSE | P15806 |
| TFE3_MOUSE.H11MO.0.A |  |
MOUSE:Tfe3 | 10 | A | 0 | dGTCACGTGA | HOCOMOCOv11 | ChIP-Seq | 0.483 | 1.0 | 2 | 30 | 478 | bHLH-ZIP factors{1.2.6} | TFE3-like factors{1.2.6.1} | 98511 | 209446 | TFE3_MOUSE | Q64092 |
| TFEB_MOUSE.H11MO.0.C |  |
MOUSE:Tfeb | 9 | C | 0 | dGTCACGTG | HOCOMOCOv9 | Integrative | 301 | bHLH-ZIP factors{1.2.6} | TFE3-like factors{1.2.6.1} | 103270 | 21425 | TFEB_MOUSE | Q9R210 | ||||
| TGIF1_MOUSE.H11MO.0.A |  |
MOUSE:Tgif1 | 9 | A | 0 | TGACAGSdv | HOCOMOCOv11 | ChIP-Seq | 0.786 | 0.979 | 2 | 5 | 137 | TALE-type homeo domain factors{3.1.4} | TGIF{3.1.4.6} | 1194497 | 21815 | TGIF1_MOUSE | P70284 |
| THA11_MOUSE.H11MO.0.B |  |
MOUSE:Thap11 | 22 | B | 0 | KGSMTbYTGGGAvdTGTAGTYY | HOCOMOCOv11 | ChIP-Seq | 0.942 | 0.784 | 2 | 2 | 128 | THAP-related factors{2.9.1} | THAP11 (HRIHFB2206){2.9.1.0.11} | 1930964 | 59016 | THA11_MOUSE | Q9JJD0 |
| THA_MOUSE.H11MO.0.C |  |
MOUSE:Thra | 17 | C | 0 | vRGKKYAnnYvAGGhCA | HOCOMOCOv11 | ChIP-Seq | 0.911 | 4 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Thyroid hormone receptors (NR1A){2.1.2.2} | 98742 | 21833 | THA_MOUSE | P63058 | ||
| THA_MOUSE.H11MO.1.C |  |
MOUSE:Thra | 9 | C | 1 | YvAGGWCAb | HOCOMOCOv11 | ChIP-Seq | 0.86 | 4 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Thyroid hormone receptors (NR1A){2.1.2.2} | 98742 | 21833 | THA_MOUSE | P63058 | ||
| THB_MOUSE.H11MO.0.D |  |
MOUSE:Thrb | 17 | D | 0 | vRGSdYAnhbnRGGhMW | HOCOMOCOv11 | ChIP-Seq | 0.784 | 4 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Thyroid hormone receptors (NR1A){2.1.2.2} | 98743 | 21834 | THB_MOUSE | P37242 | ||
| TLX1_MOUSE.H11MO.0.D |  |
MOUSE:Tlx1 | 17 | D | 0 | hYKKYAvbndvMYnRKd | HOCOMOCOv9 | Integrative | 50 | NK-related factors{3.1.2} | TLX{3.1.2.21} | 98769 | TLX1_MOUSE | P43345 | |||||
| TWST1_MOUSE.H11MO.0.B |  |
MOUSE:Twist1 | 17 | B | 0 | vCAKMTGKWhbbvATYW | HOCOMOCOv11 | ChIP-Seq | 0.969 | 0.594 | 9 | 8 | 500 | Tal-related factors{1.2.3} | Twist-like factors{1.2.3.2} | 98872 | 22160 | TWST1_MOUSE | P26687 |
| TWST1_MOUSE.H11MO.1.B |  |
MOUSE:Twist1 | 11 | B | 1 | vvCAKCTGKhh | HOCOMOCOv11 | ChIP-Seq | 0.978 | 0.604 | 9 | 8 | 470 | Tal-related factors{1.2.3} | Twist-like factors{1.2.3.2} | 98872 | 22160 | TWST1_MOUSE | P26687 |
| TYY1_MOUSE.H11MO.0.A |  |
MOUSE:Yy1 | 12 | A | 0 | vAAvATGGCSSM | HOCOMOCOv11 | ChIP-Seq | 0.984 | 0.993 | 69 | 72 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | YY1-like factors{2.3.3.9} | 99150 | 22632 | TYY1_MOUSE | Q00899 |
| UBIP1_MOUSE.H11MO.0.D |  |
MOUSE:Ubp1 | 7 | D | 0 | nCAGAdA | HOCOMOCOv9 | Integrative | 5 | CP2-related factors{6.7.2} | UBP-1 (LBP-1, SEF, CP2b){6.7.2.0.3} | 104889 | 22221 | UBIP1_MOUSE | Q811S7 | ||||
| USF1_MOUSE.H11MO.0.A |  |
MOUSE:Usf1 | 13 | A | 0 | vvGTCACGTGnbS | HOCOMOCOv11 | ChIP-Seq | 0.998 | 0.987 | 59 | 7 | 461 | bHLH-ZIP factors{1.2.6} | USF factors{1.2.6.2} | 99542 | 22278 | USF1_MOUSE | Q61069 |
| USF2_MOUSE.H11MO.0.A |  |
MOUSE:Usf2 | 19 | A | 0 | vvGTCACGTGvbvvbbvbb | HOCOMOCOv10 | ChIP-Seq | 0.998 | 0.998 | 23 | 10 | 501 | bHLH-ZIP factors{1.2.6} | USF factors{1.2.6.2} | 99961 | 22282 | USF2_MOUSE | Q64705 |
| VDR_MOUSE.H11MO.0.A |  |
MOUSE:Vdr | 16 | A | 0 | vRGKKSAbhRRGKKYR | HOCOMOCOv11 | ChIP-Seq | 0.943 | 0.932 | 48 | 30 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Vitamin D receptor (NR1I){2.1.2.4} | 103076 | 22337 | VDR_MOUSE | P48281 |
| VDR_MOUSE.H11MO.1.A |  |
MOUSE:Vdr | 9 | A | 1 | hRRGKTCAb | HOCOMOCOv11 | ChIP-Seq | 0.877 | 0.821 | 48 | 30 | 500 | Thyroid hormone receptor-related factors (NR1){2.1.2} | Vitamin D receptor (NR1I){2.1.2.4} | 103076 | 22337 | VDR_MOUSE | P48281 |
| VSX2_MOUSE.H11MO.0.C |  |
MOUSE:Vsx2 | 12 | C | 0 | hTAATTAGCYdn | HOCOMOCOv11 | ChIP-Seq | 0.996 | 3 | 504 | Paired-related HD factors{3.1.3} | VSX{3.1.3.28} | 88401 | 12677 | VSX2_MOUSE | Q61412 | ||
| WT1_MOUSE.H11MO.0.B |  |
MOUSE:Wt1 | 20 | B | 0 | vvSnGRRGGAGGRRvvRRvR | HOCOMOCOv11 | ChIP-Seq | 0.985 | 5 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | unclassified{2.3.3.0} | 98968 | WT1_MOUSE | P22561 | |||
| WT1_MOUSE.H11MO.1.A |  |
MOUSE:Wt1 | 11 | A | 1 | vdGGGAGKddv | HOCOMOCOv11 | ChIP-Seq | 0.986 | 5 | 115 | More than 3 adjacent zinc finger factors{2.3.3} | unclassified{2.3.3.0} | 98968 | WT1_MOUSE | P22561 | |||
| XBP1_MOUSE.H11MO.0.C |  |
MOUSE:Xbp1 | 12 | C | 0 | GACGTGTMhhWd | HOCOMOCOv9 | Integrative | 1910 | XBP-1-related factors{1.1.5} | XBP-1{1.1.5.0.1} | 98970 | 22433 | XBP1_MOUSE | O35426 | ||||
| ZBT17_MOUSE.H11MO.0.A |  |
MOUSE:Zbtb17 | 19 | A | 0 | vvvvSnGGRGGMGGGGvvv | HOCOMOCOv11 | ChIP-Seq | 0.797 | 0.947 | 21 | 25 | 500 | Factors with multiple dispersed zinc fingers{2.3.4} | unclassified{2.3.4.0} | 107410 | 22642 | ZBT17_MOUSE | Q60821 |
| ZBT18_MOUSE.H11MO.0.D |  |
MOUSE:Zbtb18 | 11 | D | 0 | bSCAGhTGYKb | HOCOMOCOv11 | ChIP-Seq | 0.886 | 4 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | ZNF238-like factors{2.3.3.16} | 1353609 | 30928 | ZBT18_MOUSE | Q9WUK6 | ||
| ZBT7A_MOUSE.H11MO.0.B |  |
MOUSE:Zbtb7a | 9 | B | 0 | vRGGGKCbY | HOCOMOCOv11 | ChIP-Seq | 0.929 | 12 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | ZBTB7 factors{2.3.3.8} | 1335091 | 16969 | ZBT7A_MOUSE | O88939 | ||
| ZBTB6_MOUSE.H11MO.0.D |  |
MOUSE:Zbtb6 | 13 | D | 0 | KRhSMTRGAGMYd | HOCOMOCOv11 | ChIP-Seq | 0.884 | 2 | 457 | More than 3 adjacent zinc finger factors{2.3.3} | ZBTB6-like factors{2.3.3.11} | 2442998 | 241322 | ZBTB6_MOUSE | Q8K088 | ||
| ZEB1_MOUSE.H11MO.0.B |  |
MOUSE:Zeb1 | 10 | B | 0 | bvCAGGTRdR | HOCOMOCOv11 | ChIP-Seq | 0.938 | 12 | 500 | HD-ZF factors{3.1.8} | ZEB{3.1.8.3} | 1344313 | 21417 | ZEB1_MOUSE | Q64318 | ||
| ZEP1_MOUSE.H11MO.0.D |  |
MOUSE:Hivep1 | 12 | D | 0 | KGGGAAATCCCn | HOCOMOCOv9 | Integrative | 10 | Factors with multiple dispersed zinc fingers{2.3.4} | HIV EP-factors{2.3.4.5} | 96100 | ZEP1_MOUSE | Q03172 | |||||
| ZEP2_MOUSE.H11MO.0.D |  |
MOUSE:Hivep2 | 14 | D | 0 | KSYdbGGWAASnSS | HOCOMOCOv9 | Integrative | 16 | Factors with multiple dispersed zinc fingers{2.3.4} | HIV EP-factors{2.3.4.5} | 1338076 | 15273 | ZEP2_MOUSE | Q3UHF7 | ||||
| ZFHX3_MOUSE.H11MO.0.D |  |
MOUSE:Zfhx3 | 11 | D | 0 | RTTAATWATTW | HOCOMOCOv9 | Integrative | 7 | HD-ZF factors{3.1.8} | ZFHX{3.1.8.4} | 99948 | ZFHX3_MOUSE | Q61329 | |||||
| ZFP42_MOUSE.H11MO.0.A |  |
MOUSE:Zfp42 | 12 | A | 0 | vRShGCCATYTY | HOCOMOCOv11 | ChIP-Seq | 0.909 | 0.8 | 3 | 2 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | YY1-like factors{2.3.3.9} | 99187 | 22702 | ZFP42_MOUSE | P22227 |
| ZFP57_MOUSE.H11MO.0.B |  |
MOUSE:Zfp57 | 10 | B | 0 | bYGCGGCAvv | HOCOMOCOv11 | ChIP-Seq | 0.928 | 10 | 226 | Factors with multiple dispersed zinc fingers{2.3.4} | unclassified{2.3.4.0} | 99204 | 22715 | ZFP57_MOUSE | Q8C6P8 | ||
| ZFX_MOUSE.H11MO.0.B |  |
MOUSE:Zfx | 19 | B | 0 | dSCbvGGCCTvvnvbbbbb | HOCOMOCOv9 | Integrative | 0.757 | 0.986 | 10 | 4 | 480 | More than 3 adjacent zinc finger factors{2.3.3} | ZFX/ZFY factors{2.3.3.65} | 99211 | 22764 | ZFX_MOUSE | P17012 |
| ZFX_MOUSE.H11MO.1.B |  |
MOUSE:Zfx | 8 | B | 1 | SYAGGCCb | HOCOMOCOv11 | ChIP-Seq | 0.786 | 0.991 | 10 | 4 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | ZFX/ZFY factors{2.3.3.65} | 99211 | 22764 | ZFX_MOUSE | P17012 |
| ZIC1_MOUSE.H11MO.0.B |  |
MOUSE:Zic1 | 9 | B | 0 | KGGGWGSKv | HOCOMOCOv9 | Integrative | 34 | More than 3 adjacent zinc finger factors{2.3.3} | GLI-like factors{2.3.3.1} | 106683 | 22771 | ZIC1_MOUSE | P46684 | ||||
| ZIC2_MOUSE.H11MO.0.C |  |
MOUSE:Zic2 | 15 | C | 0 | dRvYnCCTGCTGdGh | HOCOMOCOv11 | ChIP-Seq | 0.502 | 0.98 | 4 | 4 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | GLI-like factors{2.3.3.1} | 106679 | ZIC2_MOUSE | Q62520 | |
| ZIC3_MOUSE.H11MO.0.A |  |
MOUSE:Zic3 | 15 | A | 0 | dvvbhCCTGCTGdGn | HOCOMOCOv11 | ChIP-Seq | 0.943 | 5 | 170 | More than 3 adjacent zinc finger factors{2.3.3} | GLI-like factors{2.3.3.1} | 106676 | 22773 | ZIC3_MOUSE | Q62521 | ||
| ZKSC1_MOUSE.H11MO.0.A |  |
MOUSE:Zkscan1 | 19 | A | 0 | dvhdbhCCTACTRWGYGYh | HOCOMOCOv11 | ChIP-Seq | 0.961 | 6 | 500 | More than 3 adjacent zinc finger factors{2.3.3} | ZNF24-like factors{2.3.3.10} | 1921820 | 74570 | ZKSC1_MOUSE | Q8BGS3 | ||
| ZN143_MOUSE.H11MO.0.A |  |
MOUSE:Znf143 | 22 | A | 0 | dGShYbMTGGGAvdTGTAGTYY | HOCOMOCOv11 | ChIP-Seq | 0.982 | 0.979 | 28 | 9 | 463 | More than 3 adjacent zinc finger factors{2.3.3} | ZNF76-like factors{2.3.3.28} | 1277969 | 20841 | ZN143_MOUSE | O70230 |
| ZN148_MOUSE.H11MO.0.D |  |
MOUSE:Znf148 | 15 | D | 0 | dSvKbGGGGGhGGGR | HOCOMOCOv9 | Integrative | 34 | More than 3 adjacent zinc finger factors{2.3.3} | ZNF148-like factors{2.3.3.13} | 1332234 | 22661 | ZN148_MOUSE | Q61624 | ||||
| ZN281_MOUSE.H11MO.0.A |  |
MOUSE:Znf281 | 15 | A | 0 | RvRKKGGGGGAGGGG | HOCOMOCOv11 | ChIP-Seq | 0.938 | 0.999 | 7 | 3 | 502 | More than 3 adjacent zinc finger factors{2.3.3} | ZNF148-like factors{2.3.3.13} | 3029290 | 226442 | ZN281_MOUSE | Q99LI5 |
| ZN322_MOUSE.H11MO.0.B |  |
MOUSE:Znf322 | 20 | B | 0 | SdKYCTRbYMCdSdGbMKKv | HOCOMOCOv11 | ChIP-Seq | 0.888 | 0.738 | 4 | 2 | 413 | More than 3 adjacent zinc finger factors{2.3.3} | ZNF322-like factors{2.3.3.52} | 2442566 | 218100 | ZN322_MOUSE | Q8BZ89 |
| ZN335_MOUSE.H11MO.0.A |  |
MOUSE:Znf335 | 22 | A | 0 | RKSTGYCYnnnbhdvTGCCTSh | HOCOMOCOv11 | ChIP-Seq | 0.96 | 0.968 | 2 | 4 | 103 | Factors with multiple dispersed zinc fingers{2.3.4} | unclassified{2.3.4.0} | 2682313 | 329559 | ZN335_MOUSE | A2A5K6 |
| ZN335_MOUSE.H11MO.1.A |  |
MOUSE:Znf335 | 7 | A | 1 | YGCCTGW | HOCOMOCOv11 | ChIP-Seq | 0.892 | 0.995 | 2 | 4 | 123 | Factors with multiple dispersed zinc fingers{2.3.4} | unclassified{2.3.4.0} | 2682313 | 329559 | ZN335_MOUSE | A2A5K6 |
| ZN423_MOUSE.H11MO.0.D |  |
MOUSE:Znf423 | 15 | D | 0 | GGCACCCAAGGGTGC | HOCOMOCOv9 | Integrative | 22 | Factors with multiple dispersed zinc fingers{2.3.4} | ZNF423-like factors{2.3.4.12} | 1891217 | 94187 | ZN423_MOUSE | Q80TS5 | ||||
| ZN431_MOUSE.H11MO.0.C |  |
MOUSE:Znf431 | 23 | C | 0 | dvdbRvhRWCCTAAGACAGGvWb | HOCOMOCOv11 | ChIP-Seq | 0.996 | 4 | 403 | More than 3 adjacent zinc finger factors{2.3.3} | ZNF100-like factors{2.3.3.59} | 1916754 | 69504 | ZN431_MOUSE | E9QAG8 |